Characterization of Antimicrobial Resistance in Bacteria from ICU patients at a Tertiary Respiratory Care Facility in North India
Received Date: November 13, 2025 Accepted Date: December 06, 2025 Published Date: December 09, 2025
doi:10.17303/jaaa.2025.2.103
Citation: Omna Nelumbiuma, Vithal Prasad Myneedua (2025) Characterization of Antimicrobial Resistance in Bacteria from ICU patients at a Tertiary Respiratory Care Facility in North India. J Antibiot Antimicrob Agents 2: 1-11
Abstract
Purpose: Antimicrobial resistance (AMR) is a major global health concern, with increasing multidrug-resistant organisms (MDROs) in hospital settings. Local surveillance data are essential to guide antimicrobial stewardship and infection control. This study aimed to characterize bacterial pathogens and their antimicrobial susceptibility profiles among patients admitted to the respiratory intensive care unit (RICU) of a tertiary respiratory hospital in North India.
Methods: A prospective observational study was conducted in the Department of Microbiology, NITRD, New Delhi, from 23rd September 2018 to 22nd September 2019. All isolated from RICU samples submitted for bacterial culture and sensitivity were included. A total of 957 clinical specimens from 282 patients were processed using standard microbiological methods and Clinical and Laboratory Standards Institute (CLSI) guidelines. Extended spectrum -lactamase (ESBL) and metallo -lactamase (MBL) production were phenotypically detected.
Results: Of 957 samples, 312 (32.6%) yielded pathogenic bacterial growth. Non-fermenting gram-negative bacilli (NFGNB) predominated (42.3%), chiefly Acinetobacter baumannii (22.4%) and Pseudomonas aeruginosa (18.9%). Colistin showed the highest susceptibility (96.1%), followed by aminoglycosides (64%) and carbapenems (56%), while cephalosporins had poor efficacy. Among gram-positive cocci, vancomycin (100%) and linezolid (93.4%) remained highly active, with doxycycline showing good susceptibility (84.2%). ESBL and MBL production were detected in 70.2% and 37.9% of gram-negative bacilli, respectively, with 61.3% of isolates being multidrug resistant.
Conclusion: NFGNB, particularly A. baumannii and P. aeruginosa, were predominant in RICU infections. The high resistance rates underscore the urgent need for strengthened antimicrobial stewardship and rational antibiotic use.
Keywords: Antimicrobial Resistance; Respiratory ICU; Multi-Drug Resistance Organisms; Antimicrobial Stewardship
Introduction
Antimicrobial chemotherapy has been crucial in treating human infectious diseases since penicillin was discovered in the 1920s [1]. However, soon after the widespread use of antimicrobials in medicine, pathogens expressing resistance to these agents were isolated [2]. In the current era, antimicrobial resistance (AMR) has reached such a critical level that the World Health Organisation (WHO) has declared AMR a public health threat which has urged nations to formulate and implement action plans to address the issue [3].
The most recent WHO review [4, 5] about antibiotic resistance have identified a global priority pathogens list, classifying the bacteria depending on priority in critical, high and medium groups. The infections transmitted in the hospitals are majorly caused by ‘ESKAPE’ pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa and Enterobacter spp.) [6]. The National Programme on Containment of Antimicrobial Resistance [7] by National Centre for Disease Control (NCDC), New Delhi have announced a list of priority pathogens based on our country’s data which are as follows: Enterococcus spp., Staphylococcus aureus, Escherichia coli, Klebsiella spp., Acinetobacter spp., Pseudomonas spp., Salmonella enterica serotype Typhi and Salmonella enterica serotype Paratyphi.
However, the spectra of bacteria causing infections and their susceptibility pattern have been found to vary from one setting to another; a fact which highlights the importance of having local surveillance data for planning and implementing infection prevention and control (IPC) measures [8]
Objectives
The objective of the present study was to isolate and identify all aerobic and facultative anaerobic bacteria causing infections in patients admitted in the respiratory ICU (RICU) of NITRD, New Delhi from September 2018 to September 2019 and to ascertain and establish current trends of antimicrobial susceptibility profile of these isolated bacteria.
Materials and Methods
Setting
This is a prospective observational study conducted at the Department of Microbiology, NITRD, New Delhi, on all the samples for which the physician requested bacterial culture and sensitivity for the patients admitted in RICU, NITRD, New Delhi for the duration of 23rd September 2018 to 22nd September 2019.
Institutional Research committee approval was taken on 14/08/2018. Institutional Ethical Committee approval was taken on 29/09/2018. This study also was approved as part of thesis by NBE vide letter no. NBE/57/’1805042535/2018/MOD/12770.
Patient Enrolment
All patients admitted to the RICU were included in the study after duly signed informed consent. All samples received for routine bacterial culture and sensitivity were included after going through the sample acceptance and rejection criteria. Because this RICU manages only adult critically ill patients, there are no patients in the paediatric population.
Sample Collection and Processing
A total of 957 samples were collected during the study period, all of which have been recorded and analysed.
Plating of all the samples were done on blood agar and MacConkey agar. Sputum and endotracheal aspirate samples were also plated on chocolate agar and incubated overnight in candle jar. Blood samples were received in brain heart infusion (BHI) broth and incubated for 72 hours before plating on media. Urine samples were plated semi-quantitatively for determination of significance of growth.
The identified bacterial colonies were subjected to susceptibility testing against antimicrobial agents according to Clinical and Laboratory Standards Institute (CLSI) M100 document [9]. Combined disk method [9] was used to phenotypically detect extended spectrum -lactamase (ESBL) and metallo--lactamase (MBL) in gram negative bacilli. ESBL production was tested against inhibitors clavulanic acid and tazobactam using while MBL production was tested against inhibitor EDTA.
Results
A total of 282 patients were enrolled for the study from which 957 samples were sent to the laboratory. The majority of the specimens were of lower respiratory tract (LRT) (n=329, 34.38%) followed closely by urine (n=326, 34.06%) and blood (n=298, 31.14%) specimens.
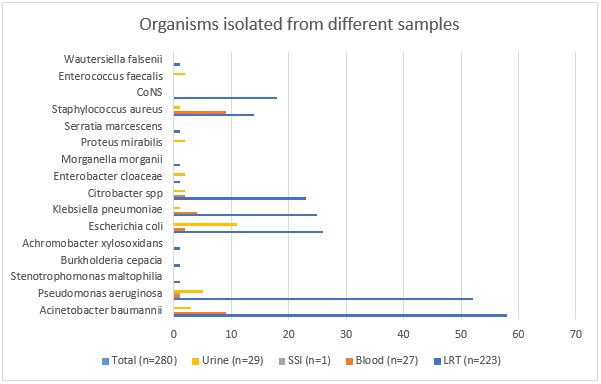
Overall, non-fermenting gram-negative bacteria (NFGNB) (n=132, 42.31%) were most commonly isolated as pathogens in cultures. Most common organism isolated from culture was Acinetobacter baumannii (n=70, 22.44%) followed by Pseudomonas aeruginosa (n=59, 18.91%).
5.1 LRT specimens: Sputum, endo-tracheal aspirate, pleural fluid and broncho-alveolar lavage were included in LRT specimens which were a total of 329 samples, 69 (20.97%) of which grew normal oropharyngeal flora, 33 (0.01%) were sterile and 3 (0.01%) samples grew contamination. Among the 223 positive pathogenic cultures, most commonly isolated organism was Acinetobacter baumannii (n=58, 26.13%) followed by Pseudomonas aeruginosa (n=52, 23.42%) as seen in Figure-1.
5.2 Blood: Blood samples and intra-vascular catheter tips were included in blood samples which came to a total of 298 samples out of which 243 (81.54%) samples were sterile on culture and 28 (9.40%) grew skin contaminants. Most commonly isolated organisms in the positive blood cultures (n=48) were Acinetobacter baumannii (n=9, 18.75%) and Staphylococcus aureus (n=9, 18.75%).
Demographic and clinical details of the patients are provided in Table-1. Antimicrobial susceptibility profile of Enterobacterales, NFGNB and gram-positive cocci (GPC) are given in Table-2. Spectrum of organisms isolated from different samples have been shown in Figure-1. Distribution of EBL-producers, MBL-producers and multi-drug resistant organisms (MDROs) have been depicted in Figure-2.
Discussion
In this study, more than half of the patients belonged to the age group 41-65 years (n=155, 54.96%) with median age of 52 years (IQR = 40-62 years) where the male patients constituted more than two-thirds of the patient population. This might be due to the fact that majority of the patients admitted in the RICU were suffering from COPD. As stated in Global Initiative for Chronic Obstructive Lung Disease (GOLD) report 2020 [10], the prevalence of COPD is appreciably higher in those ≥40 years of age as well as in men than women.
Upper respiratory tract (URT) is an area of body that is colonized early in life and stays lifelong [11]. Therefore, LRT specimens especially sputum many a times grows normal flora of the URT. In this study, the growth of normal flora was most commonly seen with LRT specimens (n=69, 20.97%). Urinary samples grew the most contamination (n=49, 15.03%), mostly from genital skin flora, probably because of lack of instructions for the proper method of collection and/or ignorance of the nursing staff during collection.
In a total of 223 positive LRT cultures, around half of the cultures reported NFGNB growth (n=113, 50.67%) with the most common organism isolated being Acinetobacter baumannii (n=58, 26.13%) followed closely by Pseudomonas aeruginosa (n=52, 23.42%). Similarly, the AMRSNAR 2019 [12] data showed that in LRT specimens, 51% of the cultures reported NFGNB group of organisms where P. aeruginosa (26%) and A. baumannii (23.4%) were grossly predominant.
Blood culture most commonly yielded Acinetobacter baumannii (n=9, 18.75%) and Staphylococcus aureus (n=9, 18.75%). In contrast, the most common organisms in blood cultures in AMRSNAR 2019[12] belonged to Enterobacteriaceae (41%) with E. coli and K. pneumoniae constituting 18% and 15% of the blood culture reports respectively. This difference might be because septicaemia in the patients at our hospital would have been caused as a complication of underlying lung infections while the AMRSNAR 2019 [12] data was collected from various reference hospitals all over the country where patients and their disease spectrum are very broad in range and involve multiple organs.
Considering gram-negative bacilli, colistin was the most effective antibiotic with an overall susceptibility of 96.06% with K. pneumoniae and E. cloaceae showing complete susceptibility and E. coli, A. baumannii and P. aeruginosa showing susceptibilities of 94-97%. With increasing use over the last five years, colistin resistance is emerging and the recent removal by CLSI [9] from susceptible category indicates that there are strains of organisms without any detectable resistance mechanism which may not respond to therapy with this drug [12].
Among the beta-lactams, cephalosporins were shown to have least susceptibility with NFGNB showing 5-13% susceptibility and Enterobacterales showed nil susceptibility. This is due to the extravagant use of the third generation cephalosporins as a common empirical therapy drug over the last three decades resulting in very high prevalence of ESBL and carbapenemase producers. It is concerning that Carbapenem susceptibility remained only average (S% = 56.03%) with A. baumannii (S% = 27.14%), Citrobacter spp (S% = 22.22%) and E. cloacae (S% = 33.33%) showing the lowest susceptibilities. However, there were organisms with higher susceptibilities like K. pneumoniae showing complete susceptibility and P. aeruginosa and E. coli showing susceptibility rates of 83.02% and 64.10% respectively. Although carbapenems have been the mainstay in empiric therapy in tertiary care ICU settings [12], regular surveillance of detection of carbapenem resistance among gram-negative bacilli should be done including molecular detection of various genes.
Aminoglycosides have showed the next best susceptibility rates (~64%) after colistin. Moreover, gentamicin had high susceptibility for not only Enterobacterales (74.75%) and NFGNB (54.26%) but also against Staphylococcus spp (81.08%) including MRSA (57.14%). However, A. baumannii showed very low Aminoglycoside susceptibility (S% = 25.71%). A marginal drop in use of aminoglycosides over the past few decades might be the reason for such high susceptibilities [12].
Gram-positive organisms showed very high susceptibility for doxycycline (84.21%) including MRSA isolates (85.71%). Therefore, doxycycline can be considered as empirical choice of therapy where gram positive infection or MRSA infection is suspected. Linezolid and vancomycin were one of the most sensitive drugs with susceptibilities of 93.42% and 100% respectively. MRSA isolates showed complete sensitivity to these drugs. Therefore, as both of them are drugs of choice for MRSA and MRCoNS, they should be reserved as a last resort.
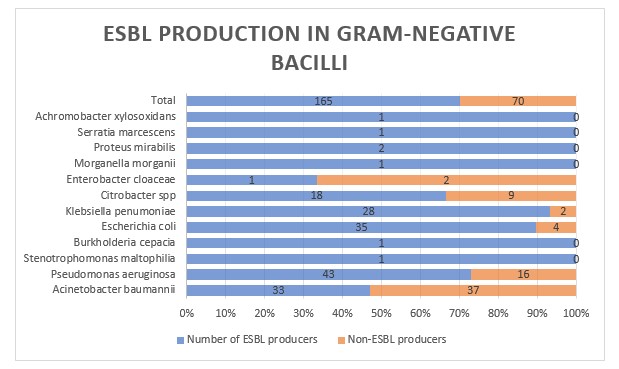
A total of 165 gram-negative bacilli (70.21%) were found phenotypically positive for ESBL production. Most of the K. pneumoniae isolates were ESBL producers (93.33%) followed closely by E. coli (89.74%) and P. aeruginosa (72.88%). The ESBL producers were significantly higher in Enterobacterales as compared to NFGNB (p = 0.00124 < 0.05). After literature search, it was found that the prevalence of ESBL was around 35-60% [13, 14] in India. A multi-centric study in the Asia-Pacific region (SMART study) [15] reported that the prevalence of ESBL rates in E. coli and K. pneumoniae respectively were 79% and 69.4%. Compared to the literature, ESBL rates were found to be quite high in our study. This might be because of the unreasonable use of third generation cephalosporins as empirical treatment in critically ill patients since past many years, with many of the patients being referred here late in the disease progression as this is a tertiary care center.
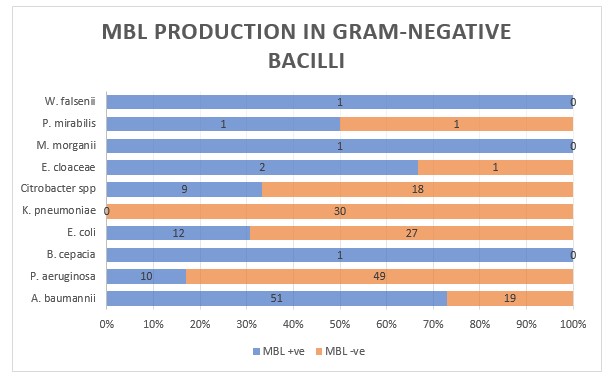
A total of 88 gram-negative bacilli (37.93%) were found phenotypically positive for MBL production. Most of the A. baumannii isolates (72.86%) were reported as MBL producers whereas none of K. pneumoniae isolates came positive for MBL. MBL production in NFGNB was reported to be significantly higher than that in Enterobacterales (p = 0.03752 < 0.05). There is confirmed unreliability of phenotypic methods of detection of ESBL and MBL production where combined disk test and double-disk synergy test have sensitivities of 92.9% and 94.9% and specificities of 96.6% and 81.4% [13]. Also, a study by Robberts F J L et al [16] reported that among E. coli isolates harbouring pAmpC genes, currently employed phenotypic ESBL detection methods perform poorly. Considering all this, the above results of ESBL and MBL rates might have false positives and negatives.
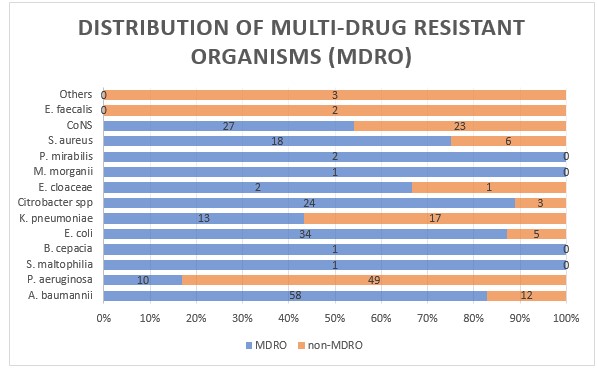
A total of 191 MDRO (61.28%) were isolated. Majority of the isolates of Citrobacter spp (88.89%), E. coli (87.18%) and A. baumannii (82.86%) were reported as multi-drug resistant organisms (MDRO). The prevalence of multi-drug resistance was found to be significantly high in Enterobacterales than in NFGNB (p = 0.00164 <0.05). In North America, a study on critically ill patients with pneumonia (DEFINE study) [17] reported a 14.1% rate of MDR infections, while a large study on nosocomial bloodstream infections conducted in 24 ICUs distributed worldwide (EUROBACT study) [18] showed on average a 47.8% MDR rate, including 20.5% and 0.5% of isolated microorganisms with extensively drug-resistant (XDR) and pan-drug-resistant (PDR) patterns, respectively. Also a study by Magira E E et al [19] on MDRO in a medical ICU found MDRO prevalence of 41.7%. The reason of our study reporting high MDRO rates might be because of unavailability of an appropriate antimicrobial stewardship in practice in our hospital. These rates necessitate more on the need of an effective stewardship program establishment for which this study has laid the foundation stone.
These results show that drugs like third generation cephalosporins and fluoroquinolones have been abused since many years for empirical therapy leading to the rise of bacterial strains that are highly resistant against these groups. Also, multi-drug resistant organisms like Stenotrophomonas maltophilia, Burkholderia cepacia and Achromobacter xylosoxidans have become emerging pathogens in an intensive care unit such that S. maltophilia has become one of the top 5 gram negative bacteria isolated from an ICU [20]. Moreover, it has become eminent that implementation of antimicrobial stewardship program in every hospital must be ensued. The tools available for the antimicrobial stewardship teams in hospitals include limiting formularies and formal restrictions of certain classes of antimicrobials, the cycling of antibiotics; and decision support, including prospective audit and feedback and, as an important component of any program, education of the prescribing staff.
Conclusion
NFGNB are the predominant pathogen causing infections in the RICU. Overall, colistin was the most susceptible drug followed by aminoglycosides and carbapenems in case of gram-negative bacilli. And among the gram-negatives, K. pneumoniae strains were more susceptible than E. coli strains and P. aeruginosa strains showed greater susceptibility than A. baumannii. Gram positive cocci showed highest susceptibilities for vancomycin and linezolid. Aminoglycosides, being old and forgotten drugs showed good susceptibility against not only gram negatives (62.66%) but also gram positives (gentamicin S% = 66.78%) including MRSA (57.14%).
Therefore, providing appropriate and regular trends of antimicrobial susceptibility to the physicians would be a good guide as to which antimicrobials to use for empirical therapy. Misuse of antibiotics should be discouraged. Turnaround time for bacterial culture and sensitivity results should be minimized as much as possible with employment of automated methods and molecular techniques for detection. And as soon as possible, empirical therapy needs to be modified with targeted therapy in accordance with the sensitivity report.
Acknowledgements
Authors sincerely thank all the staff of Department of Microbiology, NITRD, New Delhi for their constant assistance.
Conflict of Interest
The authors declare that there is no conflict of interest.
Ethical Clearance
This study was approved by the Post-Graduate Ethics Committee, National Institute of Tuberculosis and Respiratory Diseases, New Delhi, India. This study also was approved as part of thesis by NBE vide letter no. NBE/57/’1805042535/2018/MOD/12770.
- Lewis II JS, Bush K (2019) Antibacterial Agents. In: Carroll KC, Pfaller MA, Landry ML, McAdam AJ, Patel R, Richter SS, et al., editors. Manual of Clinical Microbiology. 12th ed., ASM Press; p: 1201.
- Davies JE (1997) Origins, Acquisition and Dissemination of Antibiotic Resistance Determinants. Ciba Foundation Symposium 207 - Antibiotic Resistance: Origins, Evolution, Selection and Spread, 15-35.
- Yewale VN (2014) Antimicrobial resistance - A ticking bomb! Indian Pediatr, 51: 171-2.
- Renwick MJ, Simpkin V, Mossialos E (2016) Targeting innovation in antibiotic drug discovery and development: The need for a One Health – One Europe – One World Framework [Internet]. Copenhagen (Denmark): European Observatory on Health Systems and Policies, 28806044.
- Tillotson G (2018) A crucial list of pathogens. Lancet Infect Dis, 18: 234-6.
- Miller WR, Arias CA (2024) ESKAPE pathogens: antimicrobial resistance, epidemiology, clinical impact and therapeutics. Nature Reviews Microbiology, 22: 598-616.
- National Programme on AMR Containment :: National Centre for Disease Control (NCDC) n.d. https://ncdc.gov.in/index1.php?lang=1&level=2&sublinkid=384&lid=344 (accessed April 6, 2020).
- Chaudhury A, Rani As, Kalawat U, Sumant S, Verma A, et al. (2016) Antibiotic resistance & pathogen profile in ventilator-associated pneumonia in a tertiary care hospital in India. Indian Journal of Medical Research, 144: 440.
- Clinical and Laboratory Standards Institute. CLSI M100 30th Edition. vol. 30th. 2020.
- Global Initiative for Chronic Obstructive Lung Disease. GOLD Report 2020. 2020.
- Miller JM, Binnicker MJ, Campbell S, Carroll KC, Chapin KC, et al. (2018) A Guide to Utilization of the Microbiology Laboratory for Diagnosis of Infectious Diseases: 2018 Update by the Infectious Diseases Society of America and the American Society for Microbiology. Clinical Infectious Diseases, 67: e1-94.
- Walia K, Ohri MGV, Sahni AK (2019) Annual Report Antimicrobial Resistance Surveillance and Research Network January 2019 to December 2019. 2019.
- Drieux L, Brossier F, Sougakoff W, Jarlier V (2008) Phenotypic detection of extended-spectrum β-lactamase production in Enterobacteriaceae: Review and bench guide. Clinical Microbiology and Infection, 14: 90-103.
- Kamble DMsDS (2015) Phenotypic detection of ESBL and MBL in Gram Negative bacilli isolated from clinical specimens. International Journal of Medical Research and Review, 3: 866-70.
- Lob SH, Hoban DJ, Young K, Motyl MR, Sahm DF (2018) Activity of imipenem/relebactam against Gram-negative bacilli from global ICU and non-ICU wards: SMART 2015–2016. J Glob Antimicrob Resist, 15: 12-9.
- Robberts FJL, Kohner PC, Patel R (2009) Unreliable extended-spectrum β-lactamase detection in the presence of plasmid-mediated AmpC in Escherichia coli clinical isolates. J Clin Microbiol, 47: 358-61.
- Lat I, Daley MJ, Shewale A, Pangrazzi MH, Hammond D, Olsen KM, et al. (2019) A Multicenter, Prospective, Observational Study to Determine Predictive Factors for Multidrug-Resistant Pneumonia in Critically Ill Adults: The DEFINE Study. Pharmacotherapy: The Journal of Human Pharmacology and Drug Therapy, 39: 253-60.
- Tabah A, Koulenti D, Laupland K, Misset B, Valles J, Bruzzi De Carvalho F, et al. (2012) Characteristics and determinants of outcome of hospital-acquired bloodstream infections in intensive care units: The EUROBACT International Cohort Study. Intensive Care Med, 38: 1930-45.
- Magira EE, Islam S, Niederman MS (2018) Multi-drug resistant organism infections in a medical ICU: Association to clinical features and impact upon outcome. Med Intensiva, 42: 225-34.
- Abbott IJ, Peleg AY (2015) Stenotrophomonas, achromobacter, and nonmelioid burkholderia species: Antimicrobial resistance and therapeutic strategies. Semin Respir Crit Care Med, 36: 99-110.

Tables at a glance
Figures at a glance