Successful Skin and Gut Infectious Pathogen: Staphylococccus Epidermidis, A Comprehensive Outlook of Quorum Sensing Mediated Biofilm Formation
Received Date: September 30, 2024 Accepted Date: October 30, 2024 Published Date: November 02, 2024
doi: 10.17303/jbcg.2024.6.101
Citation: Ramachandira Prabu, Govindh Boddeti, T Veeramani (2024) Successful Skin and Gut Infectious Pathogen: Staphylococccus Epidermidis, A Comprehensive Outlook of Quorum Sensing Mediated Biofilm Formation. J Bioinfo Comp Genom 6: 1-24
Abstract
Multidrug-resistant Staphylococcus epidermidis (MRSE), which forms biofilms, poses significant public health hazards through slime production and bacteremia. It induces nosocomial infections, persistent dermal damage, vaginal wall inflammation, gastrointestinal infections, endocardial infections, and gastric infections. A significant worry is biofilm development, which impedes antibiotic transport and results in a reduction in bacterial metabolic activity. Biofilms reside on plastic devices, medical prostheses, peripheral intravenous catheters, and central intravenous catheters. S. epidermidis secretes outer membrane proteins and polysaccharide intercellular adhesion (PIA) for biofilm formation, creating a conducive anaerobic environment for communal interactions with other bacteria. The microbial complex binds with extracellular matrix proteins to infiltrate the bloodstream and ultimately evade the innate immune response. This study will elucidate innovative ways for enhancing human health, focusing on immunological research, genealogy model building, candidate vaccine creation for antimicrobial-resistant strains, and the suppression of S. epidermidis biofilm production.
Keywords: Multidrug-resistant S. epidermidis; Quorum Sensing; Biofilm Formation; Biofilm Responsive Genes; Vaccine Development.
Introduction
Quorum sensing (QS) is type of behavior coordinating bacterial cell-cell interaction involved in EPS production, cell motility, toxin secretion, biofilm formation [1]. Biofilm have significance and harmful roles in industry, medical field, biomaterial implants, environment field. Nearly 40 different Staphylococcus species have been identified and more than 10 species belongs to human skin microbiome, mostly Staphylococcus epidermidis present in all humans. Microbial biofilm formation consists of typical mechanism including surface dwelling, migration, assembly, maturation, dispersal phase. Genes responsible for attachment mechanism as SesC, embp, atlE; accumulation as sesC, icaA, embp, bhp; maturation as agr, arcA and dispersal as capB and agr [2]. S. epidermidis exhibits different methodology of host attachment and accumulation, MSCRAMMs (microbial surface components recognizing adhesive matrix molecules) mediated cell-wall-anchored proteins/adhesins, hydrophobic interactions based on non-specific adhesion mechanisms, host protein-specific surface adhesion molecules [3]. S. epidermidis, almost 80% associated with biomaterial-related infection dwelling on polyurethane, polymethyl methacrylate, polyethylene, glass, teflon, polydimethylsiloxane, polyethylene terephthalate and 20% were tissue associated [6]. Propensity of S. epidermidis biofilms is greatly increases with perceptive to cell survive in toxic substances presence and decreased oxygen fluctuation level due to the available of respiratory chain-branching aids and various terminal oxidases Figure 1 [12]. Novel lantibiotics were produced by S. epidermidis multiple coagulase-negative Staphylococcus sp. colonized in the skin and synergize with cathelicidin antimicrobial peptide LL-37 act on S. aureus to inhibit growth and prevent atopic dermatitis [13]. S. aureus biofilm formation can be inhibited through S. epidermidis strains were synthesized from serine protease glutamyl endopeptidase (Esp). S. epidermidis Esp induced immune cell signaling to produce antimicrobial peptides (AMPs) from keratinocytes [13]. Small colony variants (SCVs) interlinked with persistent infection and slowly growing subpopulation were observed as predominant cause of nosocomial sepsis, however molecular mechanism of catheter mediated infection still enigmatic and ill-explored [22]. Hence need to develop novel technologies for insight into single-cell visualization of persister cell mechanics with perspective to biofilm dynamics on internal structure confirmations, biofilm stiffness, biofilm susceptible region, biofilm failure, biofilm detachment, biofilm stress management during in-vivo condition and serious pathological conditions ? Figure 4.
Genomic architecture and evolution of S. epidermidis
Complete genome sequence available from evolutionary distinct S. epidermidis strains detected in human, rodents, domesticated ruminant mammals, fermented food stuffs, meets and plants. S. epidermidis genome sequence size is about 2.5 Mb, comprises 2500 CDS, 32.2% GC content, 82 RNAs [20]. From Pan-genome studies of S. epidermidis constitutes a huge types of clonal lineages represents 20% are belongs to accessory genome in which certain genes available only in subset of isolates, whereas 80% are belongs to core genome in which genes available in all isolates [3]. Through action of bacteriophages, horizontal gene transfer (HGT) is happening frequently in S. epidermidis due to open state of pan-genome and novel clonal lineages evolution [3]. Host immune defense mechanism and antibiotics resistance genes are present due to higher number of genetic determinants with perceptive to genetically diverse of S. epidermidis [3]. Half of the genome of S. epidermidis shared with S. aureus pathogenic strains [3].
Increased genetic heterogeneity, adaptation gene, pathogenicity responsive genes, metal detoxification genes, mobile genomic elements have been transferred from HGT, which helps in colonization, survive in various host environmental condition and distinct host population [3]. Methicillin-resistant S. aureus (MRSA) and other Enterococcus spp., has developed from S. epidermidis drug resistance genes through HGT, which acting as vital reservoir [3]. Bacterial biofilm treatment requires significantly 1000 folds more concentrated antibiotics then planktonic state of similar bacteria [15]. 281 isolates were multidrug resistant S. epidermidis carried mecA gene were identified from public gathering and hospital area with frequently encountered touched surface in London [21]. 16 Multidrug resistance S. epidermidis (MRSE) studies were carried out using whole-genome sequencing approach to reveal their phylogenomic association, clonality, mobilome, virulome, resistome, pathogenicity, single-Nucleotide Polymorphism Calling [22]. The studies revealed that S. epidermidis genome carriage of multiple resistance genes contains cat(pC221), aac(6′)-aph(2′′), dfrG, erm(C), erm(B), erm(A), tet(K), blaZ and mecA conferring resistance to phenicols, aminoglycosides, macrolide–lincosamide–streptogramin B, tetracyclines, β-lactams [22]. The insertion sequence, IS256 plays critical role in S. epidermidis virulence was detected in 56.3% isolates [22]. Virulence genes shuttling and immune evasion function mediated through arginine catabolic mobile element (ACME), restriction-modification system (R-M system), bacterial anti-phage defense systems of CRISPR/CRISPR-Cas [22]. 2021). S. epidermidis virulence profile was differentiated by antiphagocytosis genes as cdsA, rmlC; immune evasion genes as manA, capC, adsA; biofilm formation responsible genes as ebp, ebh, atl, icaABDC locus [22]. Very familiar SCCmec type in S. epidermidis was belongs to community-acquired SCCmec type IV and seven MLSTs (Multilocus sequence types) were identified in CoNS S. epidermidis [22]. S. epidermidis pathogenicity is further increased due to various mobile genetic elements (MGEs) such as arginine catabolic mobile element (ACME) system for horizontal gene transfer S. epidermidis to S. aureus, conjugative transfer, phages for transmission of virulence property and resistance property acquisition, pathogenicity island for immune evasion and host colonization, transposons, insertion sequences (ISs) [24, 25]. Other factors like phenol-soluble modulins, biofilm formation, IS256 mediated genomics rearrangement, metabolic state of S. epidermidis, contributed for S. epidermidis virulence activity [24, 26]. Clinical origins and different geographic area for S. epidermidis analysis through sequence types (STs) were ST23 exhibit high resistance to rifampicin due to rpoB gene mutation, ST59, ST12 exhibit high resistance to most antibiotic drug classes, ST5 most widespread sequence types, ST2 lineage is highly dominant in hospital surroundings from Australia, clonal complex 2 [27, 28].
Horizontal gene transfer, novel variants arises by mutation and evolutionary tracking through multi-locus sequence typing (MLST) of S. epidermidis population [3]. Sequence types (STs) are two group called as ST2 and ST23 represent for diagnosed difficult bloodstream infection as well, diagnosed for adult clinical disease most like as septic arthritis, CVC infection, bacteremia [3]. Other STs called ST5, ST59, ST81, ST2 for diagnosed in neonate’s ICU, whereas ST2, ST5 for diagnosed of LOS (late-onset sepsis) [3]. One gene cluster called tarIJLM identified a mutation, which inability to form colonization in epithelial cells, however it promotes endothelial cells binding by dynamic expression of wall-teichoic acid architecture of S. epidermidis [3]. In mouse model, this mutation causes various impact on promotes bloodstream invasion and sepsis mortality [3]. ST87, ST23, ST10, ST5, ST2 on S. epidermidis isolates with increased prevalence but less number of associated infection with respect to tarIJLM mutation [3].
Clonal lineage B and A/C are two main clonal complexes or phylogenetic genotype clusters in S. epidermidis population segregated based on colonization and infection/colonization revealed from whole genome sequence studies [3, 29]. High genome plasticity presented by both clusters with exact functions of more number of genes [3]. To understand A/C lineage pathogenic potentials using pangenome-wide-association (panGWAS) comparative genomics studies [29]. A/C lineage presents higher level of antibiotics resistance, host immunity evadation, oxidative stress resistance ability leads to high level of pathogenic potential in compared to B lineage stains, which is less virulent [3]. To snapshot potentially complex and non-linear bonding between bacterial phenotype and sequence variation needs requirement of risk associated statistical models to avoid limitation of chronic infection ?
Biology of S. epidermidis infection and pathogenesis
Cutaneous microbiota plays significant role in human welfare on industrial impact, cosmetic products, aging, body odour formation, skin homeostasis [91]. Cutaneous microbiota are second largest size and diversity after intestinal microbiota of the human body [91]. Bacterial microbiota in adults such as Proteobacteria, Firmicutes, Actinobacteria are three major phyla and Staphylococcaceae, Microococcineae, Propionibacterineae, Corynebacterineae are four major species [91]. This cutaneous microbiota provides first line of barriers of skin such as furrows, hair follicles, sweat ducts, stratum corneum against pollutants, UV rays, harmful microbiota, other environmental factors [91]. Air pollutants, cosmetics are exogenous factors and natriuretic peptides, calcitonin gene related peptides, substance P are natural skin molecules affects S. aureus and S. epidermidis resistance to antibiotics, biofilm formation activity, intrinsic activity [91]. Body friction and humidity favours bacterial transfer to textiles such as polyamide, polyester are artificial fibres and cotton are natural fibers [91]. Survival of S. aureus on textiles till 3 weeks period with perceptive to composition and nature of the material [91]. Textiles controls phenomenon such as skin microorganism colonization, cutaneous microflora composition, sweat production, shin physiology, alter skin homeostasis, leads to consequence on skin cutaneous microbiota virulence, biodegradation, biofilm formation, surface alteration and finally favors to environmental opportunistic pathogens colonization on skin [91]. Flax are natural fibers and flax seed oil plays significant role on broad spectrum of antibacterial against pathogens [91]. Fax and cotton impact was investigated with perceptive to S. epidermidis and S. aureus inflammatory potential, cytotoxicity on HaCaT keratinocytes, antibiotics resistance, surface properties, biosurfactant production, biofilm formation activity, growth kinetics, however textile fibers releases stress-inducing potential and leachable with virulent toxic, due to usage of fertilizer and pesticides during plantation period [91]. S. epidermidis isolates from healthy children stool samples were infected with mice both orally and intraperitoneally for two week period. Dead mice (52.4%) were taken for histopathological examination shown that bacterial colonization in the gut causes vital organ damaged to intestine, stomach cells, spleen and kidney [7]. Neonates morbidities such as prematurity stage of retinopathy, white matter injury, bronchopulmonary dysplasia caused by S. epidermidis Figure 2 [3]. Cerebral palsy and neurodevelopmental impairment are sequalae of S. epidermidis sepsis with respective to long and short term survivors [3]. S. epidermidis is together with other skin bacterial genera Cutibacterium sp., Corynebacteium sp., Staphylococci sp. [13,15]. Pathogenicity is highly complex multifactorial scenario by combating these opportunistic bacteria needs better knowledge of the traits to developed novel candidate vaccine.
45% clinical isolates were correlates with antibiotic resistance leads to nosocomial infection and biofilm formation revealed that ica operon positive [92] . S. epidermidis is comparatively higher in the nares outside portion rather than interior of nose and more dynamics in nature of abundancy during developmental stage of humans [61]. Higher in adolescents developmental stage rather than adults and children [61]. Higher abundance in nature create an environments in nares portion against viruses (coronavirus, rhinovirus, influenza, metapneumovirus, syncytical virus) and other microbes (Proteobacteria, Cutibacterium, Moraxella, Streptococcus, Staphylococcus aureus) through prevention of respiratory tract infection, colonization and restore homeostasis in the nasal cavity, whereas low in nature leads to higher risk of infection. In the nasal epithelial cell portion of S. epidermidis may act as probiotics against respiratory pathogens as COVID-19 through interferon type I and III synthesis and represses SARS-CoV-2 (ACE2 and TMPRSS2) receptor expression [93]. Glycerol ester hydrolase (Geh), small basic protein (Sbp), Bhp, Aap, Embp involved in the abiotic and biotic surface attachment from in-vitro studies has been revealed [3]. Surface adhesions protein called S. epidermidis surface protein I (sesI) used as marker developed from in-vitro aggregation and adhension assays during initial stages of disease progression. SesI for identifying virulence and invasiveness among affected adults and neonate’s [3]. This higly prevalent in IS256, Aap protein and LOS-causing ST2 clones harboring S. epidermidis in neonates and adults, however is absent in healthy individual [3]. sdrF, sdrG, sdrH belongs to fibrinogen binding protein of ser-asp rich extracellular host matrix compound of S. epidermidis found in neonatal nasal at lower level and higher level in neonatal bloodstream isolates [3]. Rat model was infected with sdrG-positive and sdrG-deficient mutants through intravascular-catheter-associated, results in metastatic disease and bacteraemia shown in positive strain [3]. Postoperative endophthalmitis investigation conducted on adult patients with perceptive of S. epidermidis transcription profiling identified that SE1634 (staphylococcal toxin), pyruvate and iron metabolic related genes are vital contributors for pathogeneisis [3].
Combination of flavaspidic acid BB and mupirocin drug act against antibiotic-resistant Staphylococcus epidermidis mediated biofilm formation with perceptive to skin and soft tissue infections (SSTI) [94]. To understand differential gene regulation from qPCR analysis of S. epidermidis adaptation and pathogenicity at various site of the body of 11 healthy persons such as skin and nose swabs [95]. Site specific expression profile of sph gene for sphingomyelinase, SdrG, capC, dltA for immune evasion and colonization, sarA gene for regulator present in skin and nose of all individuals [95]. In case of chitinase-encoding SE0760 and agr gene (global regulator) expression was present in the skin and absent in the nose [95]. Whereas, tagB gene for wall teichoic acid (WTA) biosynthesis and sceD gene for autolysine-encoding present in nose than skin [95]. Tricarboxylic acid (TCA) cycle gene expression was downregulated and higher fold expression of sphinglomyelinase represent both anatomical niches provides better supply of nutrients [95]. Raman microscopic spectroscopy technique for intensive in situ analysis within biological systems such as proteins and small molecules spatial distribution detection, characterization and diagnosing of Staphylococcus epidermidis molecular features in human bone grafts [96]. 10 and 40 human bone graft samples have collected from infected and non-infected patients affected from S. epidermidis using Senterra II microscope [96]. Spectral resolution and principle component information have collected from reflectance data at various spectral part of S. epidermidis [96]. Major air pollutant called particulate matter (PM2.5) exposure to skin microbiota caused imbalance of various microbiome [97]. HaCaT keratinocytes exposed with particulate matter, which leads to the production of reactive oxygen species (ROS) through aryl hydrocarbon receptor (AhR) causes intrinsic mitochondrial apoptosis and mitochondrial dysfunction [97]. However, WF2R11 S. epidermidis, was treated with culture medium derived HaCaT cell lines, significantly reduced oxidative stress through AhR mediated pathway exposed to PM2.5 [97]. S. epidermidis inhibition of inflammatory cytokine secretion response through induction of ROS mediated and suppressor of AHR pathway could inhibit apoptosis and cell proliferation [97].
S. epidermidis protease act on pork myofibrillar protein (MP) converted free amino acid for enhance flavor development [98]. This protease hydrolysis of MP significantly improves stability in water, secondary structure, surface hydrophobicity identified through atomic force microscopy and Fourier transform infrared spectra [98]. HEK-293 cells shown non-cytotoxic while treated with S. epidermidis protease, however protease dependent on hydrogen bond forces act mainly bond to active site of MP [98]. S. epidermidis infection for neonatal CONS sepsis was demonstrated using preterm piglet model with perceptive to glucose metabolism and immunity for the regulation of host response during infection [99]. To support infant’s energy and growth were derived from glucose-rich parenteral nutrition, however during infection causing clinical deterioration and immune response dysregulation leads to exceed in endogenous regulation [99]. Immunoparalysis, resistance to infection and tolerance are closely connect with inflammation response, glycolysis and glucose level circulation during bacterial infection [99]. S epidermidis clinically unaffected individual revealed that provision of glucose-restricted conditions leads to moderate hypoglycemia and increase gluconeogenesis, whereas in case of clinically affected individual revealed that increase supply of parenteral glucose conditions leads to sepsis, metabolic acidosis, elevated inflammation, elevated glycolysis, hyperglycemia [99]. However, supply of normal glucose level revealed that pharmacological glycolysis inhibition and normoglycemia improves lower S. epidermidis mediated inflammation level and bacterial clearance, but unable to avoid sepsis [99]. To determine S. epidermidis protective role of spent culture fluid (SCF) and lysate (S.epi lysate) against S. aureus infected human epidermal keratinocytes model [100]. Bacterial adhesion and keratinocytes viability was evaluated from pre-exposure and post-exposure to SCF and S. epi lysate against S. aureus through reduction in adhesion and competitive displacement [100].
Bioadhesive property of K16ApoE, a cationic peptide act as anti-adhesion agent against S. epidermidis biofilm, which specifically targets on negatively charged extracellular matrix essential for bacterial entrapment and limiting therapeutics agent diffusion [101]. Breastfeeding women usually affected with mastitis caused by coagulase-negative staphylococci [102]. S. epidermidis constituted around 91% was identified from 20 women breast milk affected with mastitis were evaluated through random amplified polymorphic DNA (RAPD) and gene sequencing [102]. Sterile body fluid extracted S. epidermidis plays a clinical role in catheter-associated biofilm infection and prevalently detected Sequence type 2 (ST2) from infection site [103]. PSMδ and PSMε transcription were significantly higher in ST2 strains of body fluid may contribute for S. epidermidis dispersal leads to thicker biofilm formation [103]. Surface-exposed wall teichoic acid (WTA) polymers linked to different types of S. epidermidis clones promotes pathogenicity shift from commensal to pathogen lifestyle for infection and colonization in mice sepsis model encodes tarIJLM, accessory genetic element. S. epidermidis commensal clones were producing poly-glycerolphosphate, a type of WTA derived from multilocus sequence type 23 (MLST). HA-MRSE (Healthcare associated- methicillin resistant Staphylococcus epidermidis) clones were producing poly-ribitolphosphate (RboP-WTA), a S. aureus-type WTA carries tarIJLM, accessory genetic element responsible for pathogenesis and absent in commensal strain [104]. Rbo-WTA over-expression in mouse sepsis model caused intense endothelial attachment promotes methicillin resistance and colonization, which leads to higher mortality was observed. Currently, most of the clinical trials available for S. aureus made helpful for treating S. epidermidis at clinical level. Relatively little is known about the molecular aspects of pathogenesis and complete cure of S. epidermidis ? Figure 4.
Virulence factor responsible of S. epidermidis Infection
S. epidermidis producing soluble biofilm factor (SBF) involved in osteogenic differentiation, eventually there is no direct interaction with bacteria and mammalian cell were demonstrated from osteoblast human donors study [14]. SBF specifically inhibits phosphate and calcium settlement through regulation of RUNX2, BGLAP, COL1A1, SPP1, ALPL gene expression [14]. 6-HAP (6-N-Hydroxy aminopurine) is act as anti-tumor agent and substantial product for skin microbiome inhibit synthesis by S. epidermidis [30]. S. epidermidis commensal strains produces sphingomyelinase for host ceramides production helps to avoid skin dehydration for skin epithelial barrier homeostasis, integrity, aging and promote bacterial colonization [30]. S. epidermidis is one of key carrier function of shuttling antibiotic resistance genes (ARGs) among staphylococcal homologs group Figure 3 [31]. Catheter-related bloodstream infections (CRBSIs) of S. epidermidis contributed to 31%, which is predominant causative agents investigated in South Africa hospital were IS256 insertion element carried in the genome to 83%, mecA gene present among all isolates, β-lactams resistant [32]. S. epidermidis is skin probiotic bacteria involved in fermentation process leads to beneficial metabolites production like as short-chain fatty acids (SCFAs) from metabolize of carbon-rich molecule, support for skin disorders attenuation and act against virulent S. aureus [33]. S. epidermidis increased resistance to ultraviolet B (UV-B) and generate electricity from glycerol source [33]. Liquid coco-caprylate/caprate (LCC) promoted fermentation of S. epidermidis ATCC 12228 leads to production of short-chain fatty acids (SCFAs) production which attenuate skin disorders, act against UV-B mediated skin injuries through formation of cyclobutane pyrimidine dimers CPD) and enhance electricity generation [33].
Antibiotics, Host-immunity, Physico-chemical resistance mechanism of S. epidermidis
Underlying resistance mechanism deployed by S. epidermidis against biofilm formation remains unknown ? To address all this question, an intensive understanding of the biofilm persistence, tolerance, mechanics, and development is required. Bacterial cells getting tolerance and resistance to higher concentration of antibiotics causing gene mutation with selective survival advantage that prevent toxic effects [48]. Next stage of antibiotics tolerance through phenotypic differentiation, instead of gene mutation known as bacterial persistence and growth rejuvenation afterwards [48]. How to map S. epidermidis persister cell developed in temporal-spatial oriented biofilms and their distribution ? Figure.4 Persister cells undergo dormancy during stationary phase and revived after subculturing. What mechanism do persistent cells act upon dead cell debris after antibiotics treatment ? Figure. 4. Bacterial biofilm encounter antibiotics is temporal dependent manner, initially kills active growing cells and second phase act on sub-population level called persister cells, which prevent antibiotics and bacterial elimination occurs with least level [48]. S. epidermidis were highly resistance to hydrogen peroxide, sodium hypochlorite, sodium chloride, heat (50 °C). Screened genes were catalase, alkaline shock protein 23, σB (SigB) transcription factor dependent gene showing significantly higher expression in biofilm than planktonic cells against physico-chemical disinfection [67].
Multidrug resistance Staphylococcus sp. tolerance to antibiotics such erythromycin, kanamycin, ampicillin, fusidic acid, vancomycin, methicilin antibiotics were encoding genes mecA1, dfrC, blaZ, qacA, qacB, ccr complex [21]. σB transcription factor regulates membrane transport, cell wall antibiotics, cell wall maintenance, stationary phase entry [67, 21]. 2652 patients have been screened for microbial test from urine, hemorrhoid ulcer pus secretion, blood aerobic & anaerobic and sputum sample to understand drug resistance, antimicrobial sensitivity [67]. 1202 positive patients were segregated as 550 males and 652 females accounted for 45.76% and 54.24% among age group 70-99 years old in which 25.21% patients samples carried gram-positive bacteria mainly S. epidermidis accounted for 10.23% [67]. Multidrug-resistant S. epidermidis (MDRSE) isolates (ST2 and ST23) carries rpoB mutation confer resistance to rifampicin, teicoplanin, vancomyin, glycopeptide antibiotics identified from DNA methylation restriction modification and mutagenesis studies. [27]. Vancomycin and imipenem is crucial antibiotics for treating S. epidermidis (MRSE) methicillin-resistant strains. Imipenem is a strategic drug and broad-spectrum antibiotic against systemic infections. Imipenem resistance gene present in increased enterococci population through plasmid conjugation due to high usage of imipenem antibiotics in the last decades [55]. S. hemolyticus and S. epidermidis was reported that resistance to vancomycin antibiotics [55].
Streptomyces roseosporus derived semisynthetic lipopeptide daptomycin antibiotic which is cyclic polypeptide [68]. The native daptomycin antibiotics is anionic in nature and binding to bacterial cell membrane exhibits bactericidal activity. The unique mechanism of binding to the cell wall penetration to cytoplasmic membrane lead to rapid cell membrane depolarization causes membrane potential loss and cell death of bacteria. However, daptomycin resistance was reported in patients due to failure of treatment with respective to increased minimum inhibitory concentration (MIC) values during S. epidermidis infections [68]. 1337 patients was taken for cohort study at Ehime University Hospital during January 2013 to December 2016 with perceptive to evaluation of clinicopathological factors as sex, age, teicoplanin or vancomycin resistance and patience history of antimicrobial therapy [68]. Increased teicoplanin minimum inhibitory concentration (MIC) values with significant higher risk of daptomycin-resistance S. epidermidis in the 2.8% patients.
Prevention of PIA mediated biofilm production through DNA complex with TcaR transcriptional regulator enzyme as therapeutic strategic against infection [69]. TcaR protein structure and pharmacophore modeling are two type of drug designing methods are based on lead optimization and virtual screening of database [69]. TcaR protein structure based drug design was identified as Mol34 and 7a–7p inhibitors of S. epidermidis showed highest fitness score and binding energy with active site amino acids. Gemifloxacin drug designed from pharmacophore model generation in ZINC database using virtual screening and five TcaR novel inhibitors was identified with good binding energies score [69]. Finally, DFT simulations was performed for TcaR promising inhibitors electronic properties and adaptation of active site bioactive conformations [69]. Peptidoglycan hydrolases called lysins act against bacterial cell wall and promising approach in synergistic with antibiotics to target with biofilm and planktonic cells [18]. Exebacase is a lysin against S. epidermidis from19 clinical strains screened from prosthetic joint infections [18]. Exebacase in addition with daptomycin, vancomycin, rifampicin antibiotics shown higher anti-biomass activity [18].
S. epidermidis PSMs belongs to amphipathic peptides such as PSMβ1, PSMβ2, PSM-mec, δ-toxin (PSMγ), PSMε, PSMδ, PSMα have distinct role in interspecies competition, host immunity evasion, biofilm production, biofilm dispersion [3]. agr operon systems stringently controls S. epidermidis PSMs for low expression of virulence factors in harsh conditions [3]. S. epidermidis develops efficient defense mechanism against invading host such as efflux pumps as ABC transporters, vraFG and AMP sensor systems (aps) [3]. Rifampicin and linezolid resistance of S. epidermidis conferred through cfr (RNA methyl transferase) derived from plasmid and rpoB (RNA polymerase) derived from mutation in ribosomal gene [3]. S. epidermidis of ST23 and ST2 lineages have vancomyin/teicoplanin and rifampicin resistance derived from rpoB mutations [3]. Chlorhexidine, aminoglycosides and β-lactams antibiotics resistance was identified from prosthetic joint infections in adults revealed from comparative genomics studies of S. epidermidis with patients of commensal nasal isolates [3]. A Cationic AMP (CAMP) called melittin and synergistic effect with rifampicin and vancomycin antibiotics act against on distinct variety of gram-negative and gram-positive, mainly showing strong antibiofilm effects on methicillin-resistant, S. epidermidis [70]. Minimum biofilm eradication concentration (MBEC), minimum biofilm inhibition concentration (MBIC), Minimum biofilm preventive concentration (MBPC) was determined for testing antibiofilm with perceptive to eradication, inhibitory, preventive concentration [70]. Red Blood Cells (RBCs) and human embryonic kidney cells (HEK-293) used for analysis of melitttin activity antibiotics with respective to cytotoxicity and hemolytic activity [70]. Downregulated expression of psm, aap, icaA biofilm responsive genes with perceptive to higher therapeutic and preventive index of melittin than with other antibiotic concentration of rifampin and vancomycin [70].
S. epidermidis commensal strain from nasal epithelial cells could act as antiviral by innate immune response of interferon-related [71]. Triggers protease–protease inhibitor balance to prevented replication of influenza A virus (IAV) and favour host lungs [71]. Serpine 1, serine protease inhibitor synthesis in higher range in nasal epithelium of host induced through S. epidermidis expressing serine protease leads to inhibition of serine protease and act against IAV. Serpine 1 driven through urokinase plasminogen activator (uPA) transcription involved in human nasal epithelium to prevent IAV spread to the lungs [71]. System of CRISPR-Cas (Clustered Regularly Interspaced Short Palindromic Repeats—CRISPR associated proteins) acquires immunological infection memories provides adaptive defense strategy for prokaryotic organism [72]. Spacers called short fragments of 40 bp repetitive sequence which matches with plasmids and phages invaders in the genome and integration in the host CRISPR locus [72]. Protpacers called transcribed spacers as crRNAs (CRISPR RNAs) with guidance of Cas nucleases to target destruction of complementary nucleic acids [72]. Based on genetics constituents, six types of classification for CRISPR-Cas system [72]. In S. epidermidis CRISPR-Cas system, which is belongs to type III-A and distinct mode of spacer acquisition analyzed through inducible spacer acquisition assay [72,73]. Type III CRISPR-Cas system of S. epidermidis for increased level of spacer acquisition is different in term of spacers hotspot from tRNA and rRNA loci [72]. Minimum inhibitory concentration of Zucc essential oil (ZEO) derived from Zanthoxylum schinifolium Sieb. et Zucc, which targets cell respiratory metabolism and cell membrane for S. epidermidis bactericidal activity [75]. ZEO increases ultrastructural and morphological changes by killing S. epidermidis was detected by electron microscopy through interfering lipids membrane integrity, permeability, membrane damage and finally impaired normal physiological function [75]. Lipid acyl chains disorder, cell membrane fluidity increases and potential, increases oxidative damage through disturbs reactive oxygen species (ROS) homeostasis was analyzed through electrostatic interactions increase cell membrane depolarization and surface Zeta potential of ZEO [75].
Delftia acidovorans, human skin bacterium inhibits S. epidermidis growth through increase pH value in culture supernatant in the medium provide alkaline stress [76]. D. acidovorans produces higher ammonia for inhibition and encoding genes with perceptive to ammonia synthesis found in the genome [76]. Ammonia addition enhance the production of ROS in the culture medium of S. epidermidis, which leads to growth inhibition [76]. Sodium hydroxide was also produced with ammonia by D. acidovorans, which eventually suppresses malonic acid, a free radical scavenger involved in the inhibition of succinate dehydrogenase for N-acetyl-L-cysteine and tricarboxylic acid (TCA) cycle [76]. Moxifloxacin (MOX)-loaded liposomes for target release with optimal quantity for intraocular treatment for endophthalmitis [77]. Active loading (AL) and dehydration–rehydration (DRV) are two methods for MOX liposomes preparation for drug loading and release significantly [77]. DRV liposome was effective against S. epidermidis biofilm, growth and higher antimicrobial potential as compared with AL liposome [77]. Potent steroidal antibiotic called fusidic acid (FA) used for treatment of soft tissue and skin infection caused by Staphylococcus sp. [78]. Identified S. epidermidis fusidic acid resistance strains is prevalently found in east china [78]. FA resistant S. epidermidis was determined through agar dilution and disc diffusion method [78]. SCCmec typing, multi-locus sequence typing (MLLST), pulsed-field gel electrophoresis used for characterization of FA-resistant S. epidermidis strains [78].
Killing S. epidermidis through glucosylated liposomes consists of cholersterol, 1,2-dipalmitoyl-sn-glycero-3-phosphocholine which is natural saturated phospholipid, glucosyl moiety (GL4) which is cationic amphiphile in combination with antimicrobial trans-resveratrol (RSV) have been developed [79]. The delivering and action of RSV glucosylated liposomes by interaction with S. epidermidis cells through thin film method with the property of non-toxic, sugar moiety and cationic charge [79]. Two types of S. epidermidis cell lines was used as slime negative one (No biofilm formation) and slime positive one (Form biofilm) for evaluated RSV-loaded liposomes antimicrobial activity [79]. HPLC, electrophoretic mobility, DLS have been used for measurement of RSV entrapment concentration and efficiency, surface charge, polydispersity index, liposome mean diameter around (120–140 nm) and efficiently killing S. epidermidis at ten-fold less MIC concentration in slime positive one [79]. S. epidermidis was treated using lytic cycle inducing bacteriophages was isolated from human skin microbiome recruited for phage therapy [80]. For comparative analysis a novel phage genes were identified as N-acetylmuramoyl-L-alanine amidase for understanding interaction of S. epidermidis–phage [80].
Role of immunology in S. epidermidis pathogenesis
PIA is able to protect S. epidermidis from complement, phagocytosis, AMPs and IgG [34]. Aap, Embp, PIA prevents J774A.1 mouse macrophage mediated digestion leads to immune escape from S. epidermidis biofilm-positive [35]. Coexistence of both S. epidermidis and C. tropicalis infection in single patient to be detected for polymicrobial biofilm formation, variations in IL-6 and TNF- α are macrophage cytokine responses [36]. Platelets (PLTs) prevents S. epidermidis propagation in-vitro through downregulation of G6PD enzymes of pentose phosphate pathway responsible for glucose metabolism [37]. Successful mechanism of biofilm production in S. epidermidis to evade immunoglobulin G (IgG), C3b complement compound, host AMPs [3]. S. epidermidis metalloprotease (SepA) an extracellular proteases provide protection from neutrophil killing, complement components, host AMPs [3]. MRSE mediated PSM-mec toxins increases in survival advantage through increased cytokine expression (CXCL1, a mouse IL-8 homologue, TNF-α, IL-1β), resistance to neutrophil killing [3]. Innate immune responses raised against S. epidermidis in neonates with increased pro-inflammatory cytokines such as TNF-α, IL-8, IL-6 [3]. Immune response to neonatal sepsis with perceptive to immunosuppression and hyper inflammatory compounds involved as T cell inhibition, MDSCs and Tregs [3]. Up-regulating gene expression with perceptive to IL-6, IL-1, IFN-γ, IFN-α/β pathways in LOS preterm infants [3]. Signalling transduction of TLR/NF-κB/TREM-1 identified from transcriptional response in preterm infants cord blood monocytes inoculated with E. coli and S. epidermidis [3].
Designed 9-mer peptide vaccine against S. epidermidis through reverse vaccinology approach such as neutral metalloproteinaseas, staphylococcal secretory antigen SsaA, LysM domain protein were evaluated their non-allergenic, surface-exposed, virulent, immunogenic properties based on insilico cloning, expression, docking and simulation methodology [38]. Protein-peptide docking was performed for protein-protein interaction analysis with HLA-binding molecule of T-cell epitopes ligand and docking with Toll-like receptor (TLR-2) molecule [38]. Oral administration of Staphylococcal enterotoxin L (SELepi) and S. aureus staphylococcal enterotoxin C (SECepi) of S. epidermidis to reveal enteropathogenic properties like gut damage in Balb/c mice, may contribute in food poisoning [39]. SE-treated mice histopathological analysis from intestinal sections such as spleens, mesenteric lymph nodes, intraepithelial lymphocytes and accessed CD71 proliferation-related marker, CD69 T-cell activation marker, CD8,CD4,CD3,γδ and αβ TCR T cell lineage markers [39]. Using rat model for obtained major cell subpopulations of saprophytic S. epidermidis infection in lymph nodes [40]. This saprophytic S. epidermidis with subepidermal infection with perceptive to innate immune response of increased helper and cytotoxic T lymphocytes data was obtained cytometric evaluation [40]. Observed inflammation reduction with increased effector memory T cytotoxic lymphocytes and memory T helper lymphocytes content after three weeks later [40]. Re-infection of skin leads to immediate response of macrophages, plasmablasts, memory B lymphocytes, memory T lymphocytes, helper T lymphocytes, cytotoxic T lymphocytes [40]. Additionally number of innate immune system, MHC class II + cells, naive B lymphocytes decreased after skin re-infection, where S. epidermidis skin infection during initial and secondary time in which T regulatory lymphocyte was not seen [40]. S. epidermidis plays important role in cutaneous activation of mature IL-1β through IL-1 signalling for cytokine mediated host innate defence in human primary Keratinocytes [41]. S. epidermidis secretion of Esp serine protease caspase-1 in independent manner involved in release of mature IL-1β in Keratinocytes derived from pro‒IL-1β‒processing factor followed by proteolytic maturation [41]. H. pylori chronic infection leads to stomach cancer on the basis of risk assessment have screened H. pylori and evaluated the increased prevalence among colombian population [42]. Scenario of selection with perceptive on dietary choices, environmental toxins, H. pylori biotype and host genetic background [42]. Gastric microbiota related with non-H. pylori plays a significant role in gastric carcinogenesis [42]. Colombian patients stomach biopsy samples were screened from high-gastric-cancer-risk regions and low-gastric-cancer-risk regions [42]. Urease-positive S. epidermidis are commonly present among 59 bacterial species was identified and inoculated into INS-GAS germfree mice [42]. S. epidermidis co-infection with H. pylori leads to significantly less response of proinflammatory cytokine as IL-22, IL-17A, IL-1β than compared with mice infected with H. pylori only [42].
Role of metabolites for inhibition of S. epidermidis pathogenesis
Quercetin extract from plant inhibit S. epidermidis ATCC 35984 cell to cell adhesion by downregulation of ica locus expression and PIA synthesis [43]. Phenolic compounds derived from plant such as syringic acid and vanillin have promising against biofilm formation and maturation of methicillin-resistant S. epidermidis from three different genotypes was evaluated from EPS (extracellular polymer substance) inhibition tests, resazurin assay [44]. Inhibition of 55% EPS compounds such as polysaccharides, extracellular DNA, other proteins and inhibition of 80% biofilm formation such as genetic determinants and agr quorum-sensing systems [44]. To investigate S. epidermidis RP62A strains, how to utilize metabolic properties with perceptive to substrate auxotrophies for biomass and energy production through constructed genome-scale metabolic models [45]. Revealed that glucose is majorly utilized for biomass structure, storage and biofilm production, whereas less portion of glucose get enters into the glycolytic pathways [45]. In case of amino acids preference for biomass production and energy sources are arginine, glutamate, alanine, valine, proline [45]. In which proline, plays a critical role with perceptive to substrate auxotrophies impact on bacterial growth and metabolic models, after removal from growth media [45]. Conclusion of results with respect to collagen preference of colonization of S. epidermidis due to rich in proline amino acids [45].
Potential role of quorum sensing mediated biofilm formation in S. epidermidis
Biofilm and persister cells development is a collective bacterial behavior at extracellular matrix and sub-population level, which is essential to sustain at harsh environment remain serious challenge to human race with perspective to chronic infections ? Figure. 4. Role of human hormone atrial natriuretic peptide (ANP) inhibits S. epidermidis biofilm development, under anaerobic condition which promote aggregate dispersion and cell metabolic activity [47]. Neutrophils can only ingest pathogens of 10 μM size and biofilm range about 100 μM. Neutrophils exert pressure withstand up to 1 kPa to break biofilm into clusters and single cell, hence potentially prevent bacteria from phagocytosis [48]. Biofilm packing density divided into low, medium, high and sparse cell density represent different S. epidermidis phenotype [49]. Biofilm low cell density phenotype exhibited during vancomycin and higher salts treatment, whereas medium, high range exhibited single cluster characteristics and sparse cell type represent fractal cell clusters feature [49]. Biofilm formation is strongly stimulated by cyclic-di-GMP (cd-GMP) secondary messenger and sense by PilZ domains of BcsA at C-terminal and intracellular domain [51] (NI).
Biofilm model revealed that subinhibitory concentration of E. coli exposure to antibiotics which targets ribosome results in higher biofilm induction [54]. S. epidermidis EbpS elastin and FnbA gene expression responsible for biofilm maturity, strength and enable for infecting in-vitro and in-vivo endothelial cells [55]. EbpS gene encodes elastin-binding protein with 25 KDa in size present in the cell surface, whereas Fn (fibronectin) are soluble and immobilized form synthesis from FnbA gene [55]. S. aureus biofilm creates crucial infection in humans, however to avoid infection through interfering agent by affect cell hydrophobicity and quorum sensing receptors interference through quorum sensing inhibitors attachment. Metal ion chelator such as mn+, Zn+, Fe+ found to inhibitors of biofilm formation [56]. In S. epidermidis mature biofilm production is dependent on icaADBC operon and IS256 insertion element which is responsible for PNAG (poly-N-acetylglucosmine) synthesis, Bhp (biofilm associated-homologous protein), Embp (extracellular matrix binding protein) synthesis, Aap (accumulation-associated proteins) synthesis are the multi-step mechanisms [3]. Autolysis AltE and Aee are secreted peptidoglycan hydrolases by S. epidermidis is responsible for bacterial proliferation on cell separation and provoke medical devices primary attachment [3]. Dead cell releases eDNA (Extracellular DNA) for alternative morphotypes of S. epidermidis [3]. Quorum-sensing system of accessory gene regulator (agr) for detection of higher in S. epidermidis population density [3]. Multiple distinct agr types and agrBDCA operon involved in agr system of S. epidermidis during harsh environment downregulates key virulence genes and facilitates to undergone quiescent mode from planktonic state [3]. In preterm infants, gut is reservoir for LOS-causing (late-onset sepsis) S. epidermidis leads to bacterial dysbiosis and undeveloped intestinal mucosal barriers [3]. In preterm infants, faecal sample constitutes 90% S. epidermidis and preterm infants, meconium samples 40% S. epidermidis [3]. Patients from orthopedic device-related infection (ODRI) such as periprosthetic joint infections (PJIs) sample have been analysed and derived 111 staphylococcal strains, which belongs to S. epidermidis, multidrug resistance (MDR) and strong biofilm producing S. aureus strains [2]. This two species were analysed for genomic carriage with perceptive to virulent clinical strains in comparison with commensal strains for purpose of phenotypic and genomic characterization [2]. ST215 belongs to S. aureus and rifampicin resistance S. epidermidis associated with non/weak biofilm formation and MDR associated; whereas S. aureus and rifampicin resistance S. epidermidis belongs to both ST2 and ST45, which involved in strong biofilm production and MDR associated [2]. agr III and agr I, which is non/weak biofilm formation and strong biofilm formation of S. epidermidis MDR strain, whereas agr II of S. aureus associated with resolved infection [2].
Metalloproteinase-9, Proteinase-3, Cathepsin B, Cathepsin G called neutrophil proteases involved in provoke biofilm production of non-biofilm-synthesis S. epidermidis [57]. 11 isolates of methicillin-resistant (MR) S. epidermidis was identified from 261 samples of other MR staphylococci sp. from shrimp aquaculture farms [58]. Antibiotics such as gentamicin, trimethoprim-sulfamethoxazole and norfloxacin, erythromycin was used screening resistance genes from all staphylococcal sp. [58]. tetM, tetK , aacA-aphD, ermC, mecA are predominant antibiotics resistance gene and biofilm responsive gene was identified from all staphylococci isolates [58]. Staphylococcal Cassette Chromosome mec (SCCmec) of type V and other sequence type-spa type-SCCmec type [58]. S. epidermidis survived under harsh host environment through vital requirements for essential metals and nutrient supply was identified through regulation of fhuC and hts are two putative iron-regulated loci encodes for iron ABC transporter system involved in iron shuttling for timely associated for bacterial physiology, survival and biofilm formation [59]. Deletion analysis of single loci of either fhuC or hts results in highly affected biofilm formation of S. epidermidis and in vivo murine model showed impaired bacterial survival significantly [59]. S. epidermidis culture were infected with tibialis posterior cadaveric tendon of human to determined biofilm growth [60].
Increasing in bioburden which leads to tensile strength and elastic modulus decreased was identified from mechanical testing protocols [60]. To undermine the RT-qPCR expression and distribution of sdgB, sdgA and sdrG genes involved in in-vitro and in-vivo biofilm formation of S. epidermidis under treatment of cathepsin G [61]. 26 kDa amidase catalytic domain (AmiE) of autolysin (AtlE) surface protein is a zinc-dependent peptidoglycan peptidase revealed from NMR studies, which shed light into the interaction of S. epidermidis abiotic and biotic surface attachment with perceptive to biofilm development, bacterial colonization, cell separation, cell wall homeostasis, bacterial surface attachment and cell growth [62]. Role of S. epidermidis microvesicles and secretion was studied for biofilm formation through atmospheric scanning electron microscopy (ASEM) without and with presence of collagen [63]. Nanocarriers based Ag-decorated polymeric particles used for drug delivery system to determined S. epidermidis self-protection ability and biofilm formation at different time point from 30 min to 4 h [63]. Ag-decorated PLGA particles incubated at 30 min involved in bacterial cells adherence, damage to cell membrane and cell wall was observed through production of singlet oxygen, hydrogen peroxide, hydroxyl radicals, superoxide radicals and reactive oxygen species [63]. Whereas long incubation such as 2 h and 4 h induces drastic damage to the cells and finally collapsed, in other case Ag nanoparticles ejection was observed outside of S. epidermidis cell in the form of agglomerate [63]. Ag-decorated polymeric particles used for higher efficiency of transmission electron microscopy (TEM) analysis [63]. ASEM dishes have been prepared for analysis of biofilm formation at three distinct cultivation period as 17 h, 26 h, 30 h selected for ASEM [63]. 100 nm to 1 μm size range of microvesicles were imaged and extracellular polymeric substance was observed at 30 hrs covered the bacteria without presence of collagen, whereas nanotube-like structure was observed with presence of collagen [63]. Intercellular adhesins called Aap (accumulation-associated protein) and Embp (extracellular matrix binding protein) for accumulation and aggregation of S. epidermidis. In the absence of Aap and Embp mediated synthesis of biofilm formation in S. epidermidis showing smooth layer of bacterial accumulation. Basically, PIA synthesis induced formation of macroscopically visible, rough cell clusters, whereas Aap- and Embp-dependent biofilms preferentially displayed a smooth layer of aggregated bacteria.
IcaADBC operon role in S. epidermidis biofilm formation and pathogenesis
Guinea pig tissue model was used for comparison of wild type IcaADBC positive showing higher virulence activity than with IcaADBC mutant strains represents decreased virulence activity [64]. icaADBC transcription controlled by multitude of transcriptional regulators such as σ factor σB, SarA, TcaR and IcaR [52]. IcaR belongs to TerR family of transcriptional regulators which bind to the upstream of icaADBC promoter region and transcribed divergently [52]. IcaR binding to promoter is highly selective between icaR-icaA gene within intergenic region. sigma factor σB is global transcriptional regulatory factors involved indirectly the regulation of icaR transcription during stress response. Increased ethanol, high glucose, high NaCl represses transcription of icaR through σB involvement [52]. In S. aureus, TcaR (teicoplanin-associated locus regulator) act as repressor of icaADBC act during methicillin and teicoplanin resistance, belongs to TcaR (teicoplanin-associated locus regulator). TcaR as direct repressor for number of genes such as sarS, sasF,spa [52]. In wild type Caenorhabditis elegans, were susceptible to S. epidermidis infection in the intestine even with loss of ica locus, influence by DAF-2 insulin-signaling pathways or PMK-1 p38 mitogen-activated protein (MAP) kinase gene [65]. O-Succinyl modified dPNAG in which succinate molecules were randomly distributed conducive for Staphylococcus epidermidis clinical isolates escaping from host antibodies recognition and which in turn important targets of new therapeutics [66]. σB mutants strains correlated with higher accumulation of IcaC protein leads to higher PIA accumulation irrespective of icaADBC mRNA stability/expression [67].
PIA mediated biofilm formation
In S. epidermidis, partially deacetylated (15-20%) homopolymer of N-acetylglucosamine called PIA (polysaccharide intercellular adhesin) or PNAG (poly-N-acetylglucosamine), a cationic charged and dominant exopolysaccharide molecule. PIA or PNAG estimated molecular weight around 30 kDa encodes icaADBC operon correlated with biofilm formation, particularly required for bacterial cell accumulation and ica locus controlled by icaR, regulatory gene [74]. In S. epidermidis, 10% PIA were reported to be modified as O-succinylated rather than N-acetylated form. PS/A (capsular polysaccharide-adhesin) or PNGS (Poly-N-succinyl-b-(1–6)-glucosamine), is another type of β(1-6)-linked glucosaminyl residues of high molecular weight (>200 kDa) with modified N-succinylation (65–100%), particularly required for initial biomaterial adherence and expressed from same ica locus reported in RP62A S. epidermidis strain [74].
Composition of PS/A are galactosamine, glucosamine, galactose, uronic acids. NMR studies revealed another similar type of PIA exopolysaccharides identified in S. aureus called SAE (S. aureus exopolysaccharide) with high molecular weight greater than 300 kDa modified with O-succinylation and N-acetylation about 10% and 50-60% [74]. In Escherichia coli, Bordetella parapertussis, Yersinia pestis found similar function of ica locus called pgaABCD, bpsABCD, hmsHFRS [81-85]. Ica genes identified approximately 37% in A/C clusters and 4% in B clusters showing higher percentage towards infectious isolates rather than colonization type isolates of S. epidermidis. It was confirmed by detailed NMR analyses that N-succinylation was indeed an analytical artifact in a study that referred to S. aureus exopolysaccharide as SAE, a PIA-related molecule of high molecular weight (>300 kDa) having about 45–60% N-acetylation and 10% O-succinylation [74]. Erythrocytes hemagglutination has reported in PIA accumulation with perceptive of biomaterial-associated infections [74]. PIA mediated synthesis of biofilm formation in S. epidermidis shown rough bacteria clusters and visible by naked eye.
O-linked binding of GlcNAc to threonine and serine amino acids residues controls transcription factors and intracellular proteins such as p53, c-myc, NFκB [86]. GlcNAc provoke virulence gene expression and induce changes in morphogenesis in candida albicans, a fungal pathogen of human [87]. Changing fimbriae and CURLI fibers expression and enhance biofilm aggregation in pathogenic E. coli [87]. Production of antibiotics, changes in cell process in soil bacterial stimulated by GlcNAc [87]. GlcNAc binds to residues of fucose extracellular domains leads to ligand specific interaction is altered for notch family receptors [87]. SpoVG transcription is vital for S. epidermidis biofilm formation through modulation gene expression of icaA-icaR intergenic region for PIA production. spoVG deletion upregulating of icaR transcription gene and down regulation of icaADBC operon expression. This was observed in mice model from 1457 ΔspoVG deletion mutant and cis-complemented studies of 1457 ΔspoVG::spoVG derivative with wild type with perceptive of biofilm formation [88]. Trade-off mechanism between PIA production and antibiotic resistance in S. epidermidis [89]. Indirectly proportional association between levofloxacin, teicoplanin minimal inhibitory concentrations and formation of biofilm. PIA synthesis in S. epidermidis under growth conditions presented antibiotics as clindamycin, erythromycin, daptomycin are sensitivity to higher level, whereas sulfamethoxazole/ trimethoprim showed higher level of tolerance [89]. Prosthetic joint infection (PJI) model was evaluated from artificial synovial fluid (ASF) mimicking host surrounding and can be used as growth medium for PIA-positive and PIA-negative S. epidermidis [90]. Biofilm-positive S. epidermidis 1457 and Biofilm negative S. epidermidis mutant 1457-M10 with perceptive to PIA production and cluster formation detected in the ASF medium. embp, aap, icaA, atlE genes were upregulation, whereas agr master regulator was downregulated [90].
Omics studies in S. epidermidis pathogenesis
Comparative genomics studies were performed for screening differentially expressed genes (DEGs) from Staphylococcus epidermidis isolates in healthy conjunctiva and postoperative endophthalmitis patients by Illumina high-throughput RNA sequencing [112]. 142 pathogenesis associated genes were significantly up-regulated in genome wide transcriptional analysis from endophthalmitis strains were further validated by qRT-PCR [112]. Moreover, annotated DEGs were belongs to thioredoxin system, staphylococcal toxin (SE1634) from gene ontology and KEGG pathways revealed that two-component system, pyruvate metabolism is predominant [112]. 415 S. epidermidis isolates were classified as 141 asymptomatic carries and 274 were related with wound and bloodstream infection considered for pangenome-wide association studies. Screened 12,079 unique genes, in which 61 genes were pathogenicity-associated k-mers associated with methicillin resistance, IL-8 production, cytotoxicity, biofilm formation from accessory genome annotation. K-mers elements were identified as potential risk genotypes spread through horizontal gene transfer [113]. Comparative phylogenomics of rice endophytic S. epidermidis (RESE) which is highly distinct among 93 strains and likely to be associated with rodent strains [114]. RESE gene clusters encodes for stress tolerance and plant survival rather than for ecological adaptation represents distinct sequence identity with human S. epidermidis isolates [114]. Metagenomics-based application has unraveled difference in gene content emerged from personalized microbiota colonization to neonates can have impact on health consequence [115]. Using long-read nanopore sequencing for genome mapping of structural variants from 600 S. epidermidis strains isolated from new born. Clinically important mecA and SCCmec islands regions deletion has been identified within strain and site-specific recombination occurs at multiple sites flanked by non-canonical repeats, results in distinct pattern of antibiotic resistance and patient-specific structural variants [115]. Arginine catabolic mobile element (ACME), a genomic island may promote S. epidermidis surface colonization on human skin and In-dwelling surgical devices [116]. iIlumina sequence data taken from multidurg-resistant three isolates of genomic DNA of S. epidermidis [117]. Types of isolates are VI (4B), IV(2B&5), III(3A) holds methicillin regulators (mecR1 & mecI) and methicillin resistance gene (mecA) identified through carriage of SCCmec mobile genetic element, methicillin (mecA), Chlorhexidine (qacA), macrolide msr(A), trimethoprim (dfrG), fosfomycin (fosB), fusidic acid (fusB), penicillin (blaZ), tetracycline (tet(K), fluoroquinolone (MFS antibiotic efflux pump), aminoglycosides (aadD, aac(6′)-aph(2′) are antimicrobial resistance genes (ARGs) present in plasmids and chromosome [117]. Illumina NextSeq 500 (Illumina) was performed for isolates derived from French university hospital with respect to 16 cfr-positive linezolid-resistant S. aureus and S. epidermidis, during the period of 2015-2018 [118]. Emergence of linezolid-resistant strains of S. epidermidis isolates with plasmid carries cfr gene was received through in vivio interspecies transfer from S. aureus [118]. To determine Patrinia scabiosaefolia stress and strong antibacterial activity against MRSE from proteomic analysis using tandem mass tag-based (TMT) for screening differential expression proteins [119]. 128 proteins were up-regulated such as Hemin transport system permease (HrtB) protein, serine-aspartate repeat-containing protein C (SdrC), accumulation-associated protein (SasG), d-alanyl carrier protein (dlt), phenylalanine–tRNA ligase beta and subunit (pheT), serine–tRNA ligase (serS), carbamate kinase (arcC), ornithine carbamoyltransferase (arcB), arginine deiminase (arcA)[119]. These gene are involved in uptake of iron, biofilm formation, cell wall synthesis, protein synthesis and arginine deiminase pathway [119]. Understanding the function of genes and pathways involved in disease and health-associated abilities of S. epidermidis revealed through large scale CRISPRi (CRISPR interference) platform based on knock down studies and transcriptomics data with perceptive to growth responsive genes [120]. Selected different skin area, skin infection site and 24 different environmental conditions includes nutrient limited and multiple stress conditions in which CRISPRi, a droplet-based approach has been chosen for high-throughput functional profiling [120]. Putative essential genes for diverse environment were involved amino acid metabolism, whereas putative essential genes for survival during multiple stress conditions were involved in trace metal uptake, particularly cell wall modification in acidic stress condition [120].
Role of nanotechnology involved in S. epidermidis infection
Novel antibiotics are produced using nanotechnology by researchers now days through nanocarrier systems for low environmental risk, less manufacturing cost, minimized adverse effects and highly promising delivery. Niosomes are used for highly selective medication delivery, which is bilayer confirmation, shuttle large amount of different antimicrobial agents, water soluble in nature due to nonionic surfactants [55]. Niosome-encapsulated imipenem antibiotics have novel strategy for drug delivery system determined for MRSE antibacterial and anti-biofilm properties was examined from HDF cells with plate microtiter assay [55]. This nanocarrier was prepared from thin-film hydration method and examined for biofilm genes expression, minimum inhibitory concentration (MIC), minimum biofilm inhibitory concentration (MBIC). In different concentration of nanocarrier treatment leads to 90% cell viability. Total 162 S. epidermidis isolates were screened into 87 MRSE isolates were vancomycin-resistant and 106 methicillin-resistant [55]. Downregulated bpS, FnbA, icaD gene expression with perceptive for biofilm formation and reduce MIC and MBIC to 4 to 6 folds [55]. To determine Fe3O4 and Al2O3 nanoparticles impact on biofilm formation in S. epidermidis [128]. 10% of bacterial genes constituted for QS-control in which constitutes RNA such as sRNAs, bis- (3’-5’)-cyclic di-guanosine monophosphate (c-di-GMP), DNA, extracellular polymeric substances and remaining 97% water in biofilm environment for nutrient shuttling [128]. Fe3O4 & Al2O3 nanoparticles impact mainly on Intercellular adhesion protein C (icaC), autolysin E (atlE), extracellular matrix-binding protein (embp) shown significance gene expression level revealed in real-time quantitative polymerase chain reaction (RT-qPCR) [128].
Conclusion
Healthcare-associated and methicillin-resistant S. epidermidis (HA-MRSE) is major causative agent of infection and their molecular determinants that changes habitat of S. epidermidis remains questionable? Potential screening of virulent molecular determinants might provide way for target therapeutics. RboP-WTA is one of vital target for MRSE infections of S. epidermidis by altering growth phase leads to good therapeutic interventions. Presently uncountable methodology to combat S. epidermidis biofilms requires deep knowledge at planktonic state and group level behavior of bacterial cells? These rheological measurements could support overall knowledge about biofilm dynamics at single-cell resolution imaging. Development and optimization of animal infection model is substantial strategy for target drug delivery to various tissue specific to combat S. epidermidis biofilms and persister cell formation. Biofilm cell packing density driven mechanism from low, medium, high, sparse variation phenotypes at distinct pH and PIA concentration is still unclear. Still challengeable to discover that how S. epidermidis clonal lineages are packed within biofilms and how 3D confirmation is built within individual cells in that microbial society?
Conflicts of Interest
The authors declare no conflict of interest.
Acknowledgments
Authors are thankful to PRIST University & Sri Lakshmi Narayana Institute of Medical Sciences for providing novel ideas to carry out the review work.
- CDC (2004) National Nosocomial Infections Surveillance (NNIS) System Report. Am J Infect Control. 32: 470-85.
- Trobos M, Firdaus R, Svensson K, Tillander J, Arnellos D, Rolfson O, Thomsen P (2022) Genomics of Staphylococcus aureus and Staphylococcus epidermidis from periprosthetic joint infections and correlation to clinical outcome. Microbiol Spectr. 10: e0218121.
- Joubert IA, Otto M, Strunk T, Currie AJ (2022) Look who’s talking: host and pathogen drivers of Staphylococcus epidermidis virulence in neonatal sepsis. Int. J. Mol. Sci. 23: 860.
- Nakamura K, O’Neill AM, Williams MR, Cau L, Nakatsuji T, Horswill AR, Gallo RL (2020) Short chain fatty acids produced by Cutibacterium acnes inhibit biofilm formation by Staphylococcus epidermidis. Scientific Reports. 10: 21237.
- Salyers AA, Whitt DD (2002) Bacterial pathogenesis: a molecular approach, 2nd ed. Washington, D.C.: ASM Press. 2002. ISBN 978-1-55581-171-6.
- Gottenbos B, van der Mei HC, Busscher HJ (2000) Initial adhesion and surface growth of Staphylococcus epidermidis and Pseudomonas aeruginosa on biomedical polymers. J Biomed Mater Res. 50: 208-14.
- Akinkunmi EO, Adeyemi OI, Igbeneghu OA, Olaniyan EO, Omonisi AE, Lamikanra A (2014) The pathogenicity of Staphylococcus epidermidis on the intestinal organs of rats and mice: an experimental investigation. BMC Gastroenterol. 14: 126.
- Liu Q, Chen N, Chen H, Huang Y (2020) RNA-Seq analysis of differentially expressed genes of Staphylococcus epidermidis isolated from postoperative endophthalmitis and the healthy conjunctiva. Scientific Reports. 10: 14234.
- Brown MM, Horswill AR (2020) Staphylococcus epidermidis—Skin friend or foe? PLoS Pathog. 16: e1009026.
- Michael Otto (2009) Staphylococcus epidermidis – the “accidental” pathogen. Nat Rev Microbiol. 7: 555-67.
- Vadyvaloo V, Otto M (2005) Molecular genetics of Staphylococcus epidermidis biofilms on indwelling medical devices. Int J Artif Organs. 28:1069-78.
- Alvarezm CU, Chiquete-Fe´lix N, Uribe-Carvajal S, Penal A (2015) Exploring the Staphylococcus Epidermidis respiratory chain. Biophysical Journal. 108: 602a.
- Byrd AL, Belkaid Y, Segre JA (2018) The human skin microbiome. Nature Reviews Microbiology. 16: 143-55.
- Tubel J, Maier E, Jegen M, Marthen C, Obermeier A, Haug AT, Schneider J, Burgkart R (2021) Patient-specific effects of soluble factors from Staphylococcus aureus and Staphylococcus epidermidis biofilms on osteogenic differentiation of primary human osteoblasts. Scientific Reports. 11: 17282.
- Ceri H, Olson ME, Stremick C, Read RR, Morck D, Buret A (1999) The calgary biofilm device: new technology for rapid determination of antibiotic susceptibilities of bacterial biofilms. J. Clin. Microbiol. 37: 1771-6.
- Zhang S, Wang P, Shi X, Tan H (2021) Inhibitory properties of chinese herbal formula sanhuang decoction on biofilm formation by antibiotic-resistant Staphylococcal strains. Scientific Reports. 11: 7134.
- Ma TM, VanEpps JS, Solomon MJ (2017) Structure, mechanics, and instability of fibrin clot infected with Staphylococcus epidermidis. Biophys J. 113: 2100-9.
- Souche A, Kolenda C, Teoli J, Laurent F, Josse J (2022) Activity of Exebacase (CF-301) against biofilms formed by Staphylococcus epidermidis strains isolated from prosthetic joint infections. Antimicrob Agents Chemother. 66: e0058822.
- Yogo A, Yamamoto S, Sumiyoshi S, Iwamoto N, Aoki K, Motobayashi H, Segawa T (2022) Two cases of pyelonephritis with bacteremia by Staphylococcus epidermidis in male patients with nephrolithiasis: Case reports and a literature review. Journal of Infection and Chemotherapy. 28: 1189-92
- Raue S, Fan S, Rosenstein R, Zabel S, Luqman A, Nieselt K, Götz F (2020) The Genome of Staphylococcus epidermidis O47. Front Microbiol. 11: 2061.
- Cave R, Misra R, Chen J, Wang S, Mkrtchyan HV (2019) Whole genome sequencing revealed new molecular characteristics in multidrug resistant Staphylococci recovered from high frequency touched surfaces in London. Scientific Reports. 9: 9637.
- Liu J, Shen Z, Tang J, Huang Q, Liu Q, He L, Li M (2021) Extracellular DNA released by glycine-auxotrophic Staphylococcus epidermidis small colony variant facilitates catheter-related infections. Commun Biol. 4: 904.
- Schumacher-Perdreau F, Heilmann C, Peters G, Götz F, Pulverer G (1994) Comparative analysis of a biofilm-forming Staphylococcus epidermidis strain and its adhesion-positive, accumulation-negative mutant M7. FEMS Microbiol. Lett. 117: 71-8.
- Bouchami O, De Lencastre H, Miragaia M (2016) Impact of insertion sequences and recombination on the population structure of Staphylococcus haemolyticus. PLoS One. 11: e0156653.
- Rolo J, Worning P, Nielsen JB, Bowden R, Bouchami O, Damborg P, Miragaia M (2017) Evolutionary origin of the staphylococcal cassette chromosome mec (SCCmec). Antimicrob Agents Chemother. 61: e02302-16.
- Asante J, Hetsa BA, Amoako DG, Bester LA, Essack SY (2021) Genomic analysis of antibiotic-resistant Staphylococcus epidermidis isolates from clinical sources in the kwazulu-natal province, South Africa. Front Microbiol. 12: 656306.
- Lee JY, Monk IR, Seemann T, Kearns A, Howden BP (2018) Global spread of three multidrug-resistant lineages of Staphylococcus epidermidis. Nat. Microbiol. 3 1175-85.
- Miragaia M, Thomas J, Couto I, Enright M, De Lencastre H (2007) Inferring a population structure for Staphylococcus epidermidis from multilocus sequence typing data. J. Bacteriol. 189: 2540–52.
- Espadinha D, Sobral RG, Mendes CI, de Lencastre H, Miragaia M (2019) Distinct phenotypic and genomic signatures underlie contrasting pathogenic potential of Staphylococcus epidermidis clonal lineages. Front Microbiol. 10: 1971.
- Nakatsuji T, Fenical W, Gallo RL (2019) Response to comment on “a commensal strain of Staphylococcus epidermidis protects against skin neoplasia”. Sci Adv. 5: 5611.
- Xu Z, Misra R, Jamrozy D, Paterson GK, Cutler RR, Holmes MA (2018) Whole genome sequence and comparative genomics analysis of multi-drug resistant environmental Staphylococcus epidermidis ST59. G3: Genes Genomes Genet. 8: 2225–30.
- Ehlers MM, Strasheim W, Lowe M, Ueckermann V, Kock MM (2018) Molecular epidemiology of Staphylococcus epidermidis implicated in catheter-related bloodstream infections at an Academic Hospital in Pretoria, South Africa. Front. Microbiol. 9: 417.
- Balasubramaniam A, Adi P, Keshari S, Sankar R, Chen C, Huang C (2022) Repurposing INCI-registered compounds as skin prebiotics for probiotic Staphylococcus epidermidis against UV-B. Scientific Reports. 10: 21585.
- Chiquet C , Musson C, Aptel F, Boisset S, Maurin M (2018) Genetic and phenotypic traits of Staphylococcus epidermidis strains causing postcataract endophthalmitis compared to commensal conjunctival flora. Am J Ophthalmol. 191: 76-82.
- Schommer NN, Christner M, Hentschke M, Ruckdeschel K, Aepfelbacher M, Rohde H (2011) Staphylococcus epidermidis uses distinct mechanisms of biofilm formation to interfere with phagocytosis and activation of mouse macrophage-like cells 774A.1. Infect Immun. 79: 2267-76.
- Phuengmaung P, Panpetch W, Singkham-In U, Chatsuwan T, Chirathaworn C, Leelahavanichkul A (2021) Presence of Candida tropicalis on Staphylococcus epidermidis biofilms facilitated biofilm production and Candida dissemination: an impact of fungi on bacterial biofilms. Front. Cell. Infect. Microbiol. 11: 763239.
- Li Z, An Q, Xu J, Xin j, Chen Y, An N, Gu S, Yi J, Yin W (2022) Platelets inhibit the proliferation of Staphylococcus epidermidis by directly down-regulating G6PD. Blood & Genomics, 6: 28-35.
- Sethi G, Sethi S, Krishna R (2022) Multi-epitope based vaccine design against Staphylococcus epidermidis: A subtractive proteomics and immunoinformatics approach. Microbial Pathogenesis. 165: 105484.
- Tabis A, Gonet M, Schubert J, Miazek A, Nowak M, Tomaszekv A, Bania J. Analysis of enterotoxigenic effect of Staphylococcus aureus and Staphylococcus epidermidis enterotoxins C and L on mice. Microbiological Research; 2022. 258,126979.
- Cąkała-Jakimowicz M, Puzianowska-Kuznicka M. Towards understanding the lymph node response to skin infection with saprophytic Staphylococcus epidermidis. Biomedicines 2022, 10 (5), 1021.
- Rademacher F, Bartels J, Glaser R, Rodewald M, Schubert S, Drucke D, Rohde H, Harder J (2022) Staphylococcus epidermidis-derived protease Esp mediates proteolytic activation of pro‒IL-1β in human keratinocytes. Journal of Investigative Dermatology. 2022; S0022-202X (22)00306-2.
- Shen Z, Dzink-Fox J, Feng Y, Muthupalani S, Peek RM, Wang TC, Fox JG (2022) Gastric non-Helicobacter pylori urease-positive Staphylococcus epidermidis and Streptococcus salivarius isolated from humans have contrasting effects on H. pylori-associated gastric pathology and host immune responses in a murine model of gastric cancer. ASM Journals. 7: e0077221.
- Mu Y, Zeng H, Chen W (2021) Quercetin inhibits biofilm formation by decreasing the production of EPS and altering the composition of EPS in Staphylococcus epidermidis. Front. Microbiol. 12: 631058.
- Minich A, Levarski AZ, Liptakova D, Stuchlik S (2022) Complex analysis of vanillin and syringic acid as natural antimicrobial agents against Staphylococcus epidermidis biofilms. Int. J. Mol. Sci. 23: 1816.
- Calvo TD, Tejera N, McNamara I, Langridge GC, Wain J, Poolman M, Singh D (2022) Genome-scale metabolic modelling approach to understand the metabolism of the opportunistic human pathogen Staphylococcus epidermidis RP62A. Metabolites. 12: 136.
- Hall-Stoodley L., Costerton JW, Stoodley P (2004) Bacterial biofilms: from the natural environment to infectious diseases. Nat. Rev. Microbiol. 2: 95-108.
- Ovcharova MA, Geraskina OV, Plakunov VK, Gannesen AV (2021) Atrial natriuretic peptide affects skin commensal Staphylococcus epidermidis and Cutibacterium acnes dual-species biofilms. Microorganisms. 9: 552.
- Yan J, Bassler BL (2019) Surviving as a community: antibiotic tolerance and persistence in bacterial biofilms. Cell Host Microbe. 26: 15-21.
- Stewart EJ, Satorius AE, Younger JG, Solomon MJ (2013) Role of environmental and antibiotic stress on Staphylococcus epidermidis biofilm microstructure. Langmuir. 29: 7017-24.
- Jager S, Mack D, Rohde H, Horstkotte MA, Knobloch JK (2005) Disintegration of Staphylococcus epidermidis biofilms under glucose-limiting conditions depends on the activity of the alternative sigma factor sigma B. Appl. Environ. Microbiol. 71: 5577–81.
- Morgan JLW, Strumillo J, Zimmer J (2013) Crystallographic snapshot of cellulose synthesis and membrane translocation. Nature. 493: 181-6.
- Hoang T, Zhou C, Lindgren JK, Galac MR, Corey B. Olson ME, Fey PD (2019) Transcriptional Regulation of icaADBC by both IcaR and TcaR in Staphylococcus epidermidis. J Bacteriol. 201: e00524-18.
- Martinez-Garcia S, Betanzos‐Cabrera G, Jan‐Roblero J, Rodríguez‐Martínez S, Cancino‐Diaz ME, Cancino‐Diaz JC (2019) Non-biofilm-forming commensal Staphylococcus epidermidis isolates produce biofilm in the presence of trypsin. Microbiology open. 8: e906.
- Boehm A, Steiner S, Zaehringer F, Casanova A, Hamburger F, Ritz D, Keck W (2009) Ackermann M, Schirmer T, Jenal U. Second messenger signalling governs Escherichia coli biofilm induction upon ribosomal stress. Mol. Microbiol. 72: 1500-16.
- Piri-Gharaghie T, Jegargoshe-Shirin N, Saremi-Nouri S, Hoseinnezhad-lazarjani E, Mousavi A, Farhadi-Biregani A, Fatehi-Ghahfarokhi S (2022) Effects of Imipenem-containing Niosome nanoparticles against high prevalence methicillin-resistant Staphylococcus Epidermidis biofilm formed. Scientific Reports. 12: 5140.
- Ahmad N; Shiva J; Taher A; Leila N (2022) Staphylococcus aureus: biofilm formation and strategies against it. Current Pharmaceutical Biotechnology. 23: 664-78.
- Gomez-Alonso IS, Martínez-Garcia S, Cancino-Diaz ME, Cancino J, Cancino-Diaz JC (2022) Low concentration of the neutrophil proteases cathepsin G, cathepsin B, proteinase-3 and metalloproteinase-9 induce biofilm formation in non-biofilm-forming Staphylococcus epidermidis isolates. Int. J. Mol. Sci. 23: 4992.
- Rajan V, Sivaraman GK, Vijayan A, Elangovan R, Prendiville A, Bachmann TT (2022) Genotypes and phenotypes of methicillin-resistant staphylococci isolated from shrimp aquaculture farms. Environ Microbiol Rep. 14: 391-9.
- Oliveira F , Lima T, Correia A, Silva AM, Soares C, Morais S, Weißelberg S, Vilanova M, Rohde H, Cerca N (2022) Involvement of the iron-regulated loci hts and fhuC in biofilm formation and survival of Staphylococcus epidermidis within the host. Microbiology Spectrum. 10: e02168-21.
- Sorensen HH, Magnussen RA, DiBartola AC, Mallory NT, Litsky AS, P, Swinehart SD, Kaeding CC, Flanigan DC (20222) Influence of Staphylococcus epidermidis biofilm on the mechanical strength of soft tissue allograft. J Orthop Res.
- Gómez-Alonso IS, Chavez-Cabrera C, Rodríguez-Martínez S, Cancino-Diaz ME, Cancino-Diaz JC (2022) The expression of glycosyltransferases sdgA and sdgB in Staphylococcus epidermidis depends on the conditions of biofilm formation. Archives of Microbiology. 204: 274.
- Yadav R, Somarathne RP, Yang S, Youn PJ, Fitzkee NC (2022) NMR illuminates the ligand and surface adsorption of autolysin-amidase from Staphylococus epidermidis. Biophysical journal 121.
- Takahashi C, Moriguchi K, Hori M, Kawai T, Sato M, Sato C. Biofilm formation of Staphylococcus epidermidis with and without collagen imaged using atmospheric scanning electron microscopy and antibacterial effect of Ag-decorated polymeric particles imaged by transmission electron microscopy. Microsc. Microanal. 2022; 28.
- Chokr A, Leterme D, Watier D, Jabbouri S (2007) Neither the presence of ica locus, nor in vitro-biofilm formation ability is a crucial parameter for some Staphylococcus epidermidis strains to maintain an infection in a guinea pig tissue cage model. Microb. Pathog. 42: 94–7.
- Begun J, Gaiani JM, Sifri CD (2007) Staphylococcal biofilm exopolysaccharide protects against Caenorhabditis elegans immune defenses. PLoS Pathog. 3: e57.
- Gening ML, Litran TM, Kropec A, Skurnik D, Grout M, Tsvetkov YE, Nifantiev NE, Pier GB (2010) Synthetic beta-(1->6)-linked N-acetylated and nonacetylated oligoglucosamines used to produce conjugate vaccines for bacterial pathogens. Infect. Immun. 78: 764-72.
- Valle J, Echeverz M, Lasa I (2019) σB Inhibits Poly- N-acetylglucosamine exopolysaccharide synthesis and biofilm formation in Staphylococcus aureus. J Bacteriol. 201: e00098-19.
- Watanabe S, Kawakami Y, Kimura H, Murakami S, Miyamoto H, Takatori S, Maki J, Araki H, Yamaguchi T (2019) Association between daptomycin susceptibility and teicoplanin resistance in Staphylococcus epidermidis. Scientific Reports. 9: 18533.
- Vuppala S, Kim J, Choi J, Jang J (2022) A combination of pharmacophore-based virtual screening, structure-based lead optimization, and DFT Study for the identification of S. epidermidis TcaR inhibitors. Pharmaceuticals. 15: 635.
- Mirzaei R, Alikhani MY, Arciola CR, Yousefimashouf R, Bagheri KP (2022) Prevention, inhibition, and degradation effects of melittin alone and in combination with vancomycin and rifampin against strong biofilm producer strains of methicillin-resistant Staphylococcus epidermidis. Biomedicine Pharmacotherapy. 147: 112670.
- Jo A, Won J, Gil CH, Kim SK, Lee K, Yoon SS, Kim HJ (2022) Nasal symbiont Staphylococcus epidermidis restricts the cellular entry of influenza virus into the nasal epithelium. npj Biofilms and Microbiomes. 8: 26.
- Aviram N, Thornal AN, Zeevi D, Marraffini LA (2022) Different modes of spacer acquisition by the Staphylococcus epidermidis type III-A CRISPR-Cas system. Nucleic Acids Research. 50: 3.
- Pyenson NC, Gayvert K, Varble A, Elemento O, Marraffini LA (2017) Broad targeting specificity during bacterial type III CRISPR-Cas immunity constrains viral escape. Cell Host Microbe. 22: 343-53.
- Maira-Litrán T, Kropec A, Abeygunawardana C, Joyce J, Goldmann DA, Pier GB (2002) Immunochemical properties of the staphylococcal Poly-N-Acetylglucosamine surface polysaccharide. Infect Immun. 70: 4433–40.
- Liao S, Yang G, Ou Y, Huang S, Li B, Li A, Kan J (2022) Inhibitory impacts of essential oil (Zanthoxylum schinifolium Sieb. et Zucc) on the growth of Staphylococcus epidermidis. Food Bioscience. 49: 101906.
- Matsumoto Y, Ogasawara Y, Sugita T, Ohkubo T (2022) Alkaline stress inhibits the growth of Staphylococcus epidermidis by inducing TCA cycle-triggered ROS production. Biochemical and Biophysical Research Communications. 588: 104-10.
- Natsaridis E, Gkartziou F, Mourtas S, Spiliopoulou I, Antimisiaris SG (2022) Moxifloxacin liposomes: effect of liposome preparation method on physicochemical properties and antimicrobial activity against Staphylococcus epidermidis. Pharmaceutics. 14: 370.
- Rao L, Chen S, Lin C (2022) The dissemination of fusidic acid resistance among Staphylococcus epidermidis clinical isolates in Wenzhou, China. Infect Drug Resist. 15: 2537–44.
- Bombelli C, Pagano L, Spiliopoulou I, Sophia G. Antimisiaris SC, Mancini G (2022) Resveratrol loaded in cationic glucosylated liposomes to treat Staphylococcus epidermidis infections. Chemistry and Physics of Lipids. 243: 105174.
- Valente LG, Pitton M, Furholz M, Oberhaensli S, Bruggmann R, Leib SL, Jakob SM, Cameron DR (2021) Isolation and characterization of bacteriophages from the human skin microbiome that infect Staphylococcus epidermidis. FEMS Microbes, 2: xtab003.
- Bobrov AG, Kirillina Olga, Forman S, Mack D, Perry RD (2008) Insights into Yersinia pestis biofilm development: topology and co-interaction of Hms inner membrane proteins involved in exopolysaccharide production. Environ Microbiol. 10: 1419–32.
- Itoh Y, Romeo T (2005) Depolymerization of beta-1,6-N-acetyl-D-glucosamine disrupts the integrity of diverse bacterial biofilms. J Bacteriol. 187: 382–7.
- Parise G, Mishra M (2007) Role of a putative polysaccharide locus in Bordetella biofilm development. J Bacteriol. 189: 750–60.
- Ganguly T, Johnson JB (2014) The Bordetella pertussis Bps polysaccharide enhances lung colonization by conferring protection from complement-mediated killing. Cell Microbiol. 16: 1105–18.
- Nicholson TL, Deora R (2017) The bordetella bps polysaccharide is required for biofilm formation and enhances survival in the lower respiratory tract of swine. Infect Immun. 85.
- Özcan S, Andrali SS, Cantrell JEL (2010) Modulation of transcription factor function by O-GlcNAc modification. Biochim Biophys Acta. 1799: 353–64.
- Konopka JB (2012) N-Acetylglucosamine Functions in Cell Signaling. Scientifica. 2012; Article ID 489208: 15
- Benthien H, Fresenborg B, Molle V, Knobloch JK, Bischoff M (2022) The transcription factor SpoVG is of major importance for biofilm formation of Staphylococcus epidermidis under In vitro conditions, but dispensable for In vivo biofilm formation. Int. J. Mol. Sci. 23: 3255.
- Carcione D, Leccese G, Landini P, Brilli M, Paroni M (2022) Lack of direct correlation between biofilm formation and antimicrobial resistance in clinical Staphylococcus epidermidis isolates from an italian hospital. Microorganisms. 10: 1163.
- Stamm J, Weißelberg S , Both A, Failla AV, Nordholt G, Büttner H, Aepfelbacher M, Rohde H (2022) Development of an artificial synovial fluid useful for studying Staphylococcus epidermidis joint infections. Front Cell Infect Microbiol. 12: 948151.
- Catovic C, Abbes I, Barreau M, Chevalier S, Feuilloley MGJ (2022) Cotton and flax textiles leachables impact differently cutaneous Staphylococcus aureus and Staphylococcus epidermidis biofilm formation and cytotoxicity. Life. 12: 535.
- Cafiso V, Bertuccio T, Santagati M, Campanile F, Amicosante G, Perilli MG, Stefani S (2004) Presence of the ica operon in clinical isolates of Staphylococcus epidermidis and its role in biofilm production. Clinical Microbiology and Infection. 10: 1081-8
- Ortega-Peña S, Rodríguez-Martínez S, Cancino-Diaz ME, Cancino-Diaz JC (2022) Staphylococcus epidermidis controls opportunistic pathogens in the nose, could it help to regulate SARS-CoV-2 (COVID-19) infection? Life (Basel). 12: 341.
- Zhiling C, Zitong M, Shiqian Z, Shihua L, Shengjun X, Jinghui L, Chunping T, Zhibin S (2022) Flavaspidic acid BB combined with mupirocin improves its anti-bacterial and anti-biofilm activities against Staphylococcus epidermidis. BMC Microbiology. 22: 179.
- Teichmann P, Both A, Wolz C. Burian M (2022) The Staphylococcus epidermidis transcriptional profile during carriage. Front. Microbiol. 13: 896311.
- Wurm Putzer D, Kappacher C, Huck CW, Pallua JD (2022) Raman microscopic spectroscopy as a diagnostic tool to detect Staphylococcus epidermidis in bone grafts. Spectrochimica Acta Part A: Molecular and Biomolecular Spectroscopy. 280: 121570.
- Lee E, Ahn H, Shinyoung Park, Gihyeon Kim, Park H (2022) Staphylococcus epidermidis WF2R11 Suppresses PM2.5-Mediated Activation of the Aryl Hydrocarbon Receptor in HaCaT Keratinocytes. Probiotics and Antimicrobial Proteins. 14: 915–33.
- Wang H, Xu J, Liu Q, Chen Q, Sun F, Kong B (2022) Interaction between protease from Staphylococcus epidermidis and pork myofibrillar protein: Flavor and molecular simulation. Food Chemistry. 386: 132830.
- Muk T, Brunse A, Henriksen NL, Aasmul-Olsen K, Nguyen DN (2022) Glucose supply and glycolysis inhibition shape the clinical fate of Staphylococcus epidermidis–infected preterm newborns. JCI insight. 7: e157234.
- Mohammedsaeed W, Manzoor N (2022) An in vitro investigation of the protective role of Staphylococcus epidermidis extracts on Staphylococcus aureus induced toxicity in human keratinocytes. Indian Journal of Medical Microbiology. 40: 239-44.
- Ahlschwede KM, Amissah F, Deshmukh R (2022) Evaluating the effect of cationic peptide K16ApoE against Staphylococcus epidermidis biofilms. Journal of Pharmaceutical Investigation. 52: 139–49.
- Cullinane M, Scofield L, Murray GL, Payne MS, Amir LH (2022) Random amplified polymorphic DNA analysis reveals no clear link between Staphylococcus epidermidis and acute mastitis. Anzjog. 62: 605-9.
- Jin Y, Wang Q, Li M, Liu Q (2022) PSM contributes to the dispersal of Staphylococcus epidermidis isolates from catheters. Front Microbiol. 13: 934358.
- Rohde H, Winstel V, Peschel A (2021) Staphylococcus epidermidis clones express Staphylococcus aureustype wall teichoic acid to shift from a commensal to pathogen lifestyle. Nature Microbiology. 6:757–68.
- Bartel DP (2018) Metazoan microRNAs. Cell. 173: 20–51.
- Aguilar C, Mano M, Eulalio A (2019) MicroRNAs at the host–bacteria interface: host defense or bacterial offense. Trends in Microbiology. 27: 206-18.
- Taganov KD, Boldin MP, Chang KJ, Baltimore D (2006) NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA. 103: 12481–6.
- Liu PT, Wheelwright M (2012) MicroRNA-21 targets the vitamin D-dependent antimicrobial pathway in leprosy. Nat. Med. 18: 267–73.
- Teng GG, Wang W, Wang S, Chu Y, Li J, Dai Y (2013) Let-7b is involved in the inflammation and immune responses associated with Helicobacter pylori infection by targeting Toll-like receptor 4. PLoS One. 8: e56709.
- Matsushima K. Isomoto H (2011) MicroRNA signatures in Helicobacter pylori-infected gastric mucosa. Int. J. Cancer. 128: 361–70.
- Pagliari M. Cagnin S (2017) Helicobacter pylori affects the antigen presentation activity of macrophages modulating the expression of the immune receptor CD300E through miR-4270. Front. Immunol. 8: 1288.
- Liu Q, Chen N, Chen H, Huang Y (2020) RNA-Seq analysis of differentially expressed genes of Staphylococcus epidermidis isolated from postoperative endophthalmitis and the healthy conjunctiva. Scientific Reports. 10: 14234.
- Mack D, Corander J, Sheppard SK (2018) Disease-associated genotypes of the commensal skin bacterium Staphylococcus epidermidis. Nature Communications. 9: 5034.
- Chaudhry V, Patil PB (2016) Genomic investigation reveals evolution and lifestyle adaptation of endophytic Staphylococcus epidermidis. Scientific Reports. 6: 19263.
- Borenstein-Levin L, Kugelman A, Kishony R (2021) Rapid methicillin resistance diversification in Staphylococcus epidermidis colonizing human neonates. Nature Communications, 12: 6062.
- Jin Y, Chen Z, Yu F (2019) Antimicrobial susceptibility, virulence determinants profiles and molecular characteristics of Staphylococcus epidermidis isolates in Wenzhou, eastern China. BMC Microbiology. 19: 157.
- Altayb HN, Elbadawi HS, Baothman O, Hosawi S, Chaieb K (2022) Whole-Genome sequence of multidrug-resistant methicillin-resistant Staphylococcus epidermidis carrying biofilm-associated genes and a unique composite of SCCmec. Antibiotics. 11: 861.
- Figueiredo AMS, Dupieux C (2022) Persistence of a multidrug-resistant worldwide-disseminated methicillin-resistant Staphylococcus epidermidis clone harboring the cfr linezolid resistance gene in a French hospital with evidence of interspecies transfer to several Staphylococcus aureus lineages. Journal of Antimicrobial Chemotherapy. 77: 1838–46.
- Liu X, An L, Ren S, Zhou Y, Peng W (2022) Comparative proteomic analysis reveals antibacterial mechanism of Patrinia scabiosaefolia against methicillin resistant Staphylococcus epidermidis. Infect Drug Resist. 15: 883–93.
- Spoto M, Guan C, Nzutchi YO, Kim D, Oh J (2022) Large-scale CRISPRi and transcriptomics of Staphylococcus epidermidis identify genetic factors implicated in commensal-pathogen lifestyle versatility. Biorxiv. 15: 883-93.
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J. Mol. Biol. 215: 403–10.
- Joensen KG, Scheutz F, Lund O, Hasman H, Kaas RS, Nielsen EM (2014) Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J. Clin. Microbiol. 52: 1501–10.
- Larsen MV, Cosentino S, Rasmussen S, Friis C, Hasman H, Marvig RL (2012) Multilocus sequence typing of total-genome-sequenced bacteria. J. Clin. Microbiol. 50: 1355–61.
- Thomas JC, Vargas MR, Miragaia M, Peacock SJ, Archer GL, Enright MC (2007) Improved multilocus sequence typing scheme for Staphylococcus epidermidis. J. Clin. Microbiol. 45: 616-19.
- Feil EJ, Li BC, Aanensen DM, Hanage WP, Spratt BG (2004) eBURST: inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. J. Bacteriol. 186: 1518–30.
- Aziz RK, Bartels D, Best AA, Dejongh M, Disz T, Edwards RA, Zagnitko O, Overbeek RA (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genomics. 9: 75.
- Arndt D, Grant JR, Marcu A, Sajed T, Pon A, Liang Y, Wishart DS (2016) PHASTER: a better, faster version of the PHAST phage search tool. Nucleic Acids Res. 44 W16–21.
- Mahdi SA, Al-Musawi S (2022) Impact of Al2O3 & Fe3O4 nanoparticles on genes expression levels on biofilm in Staphylococcus epidermidis. AIP Conference Proceedings. 2437: 020031.

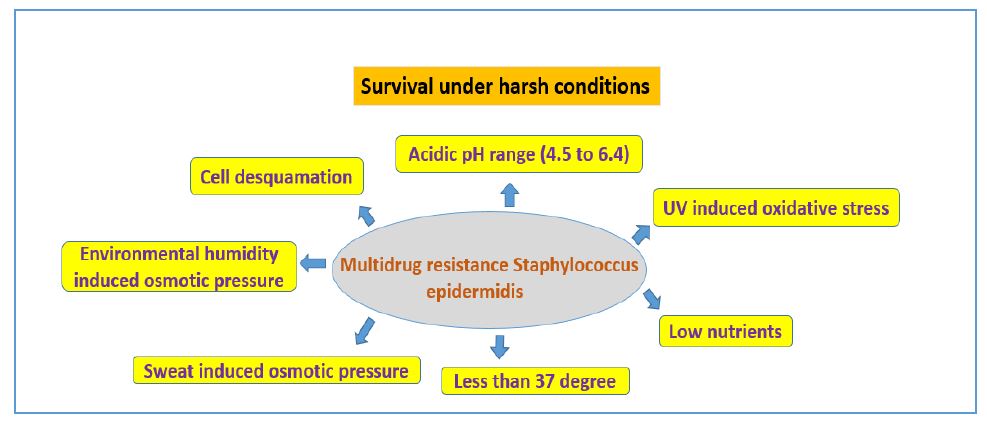
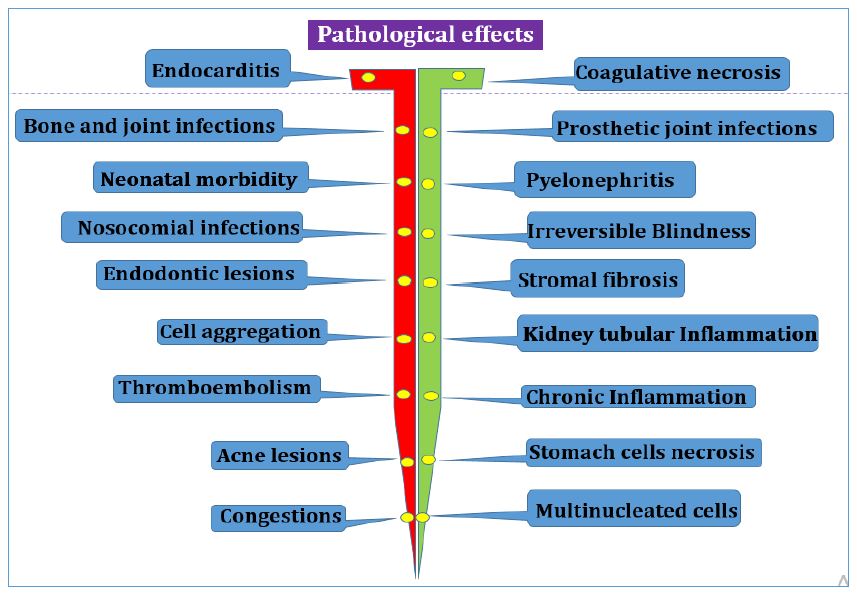
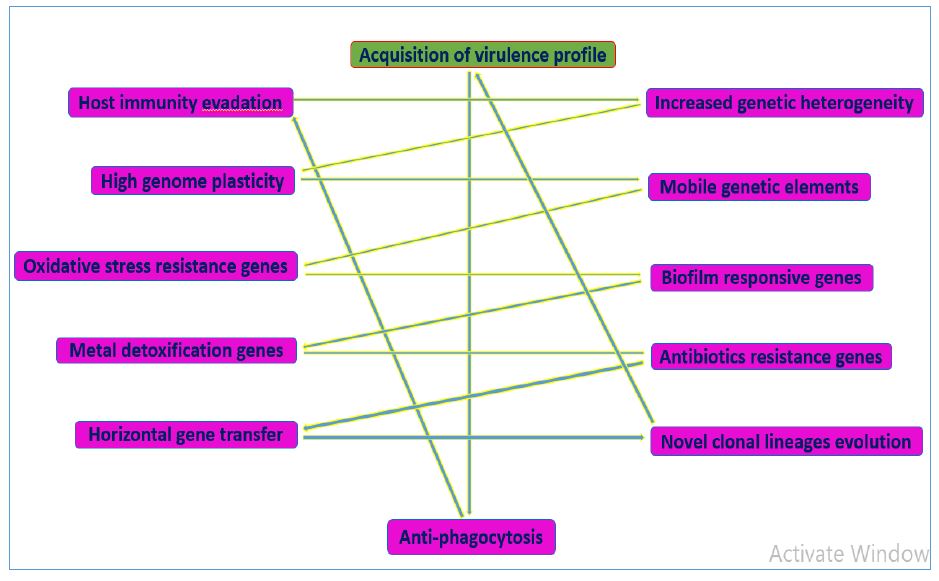
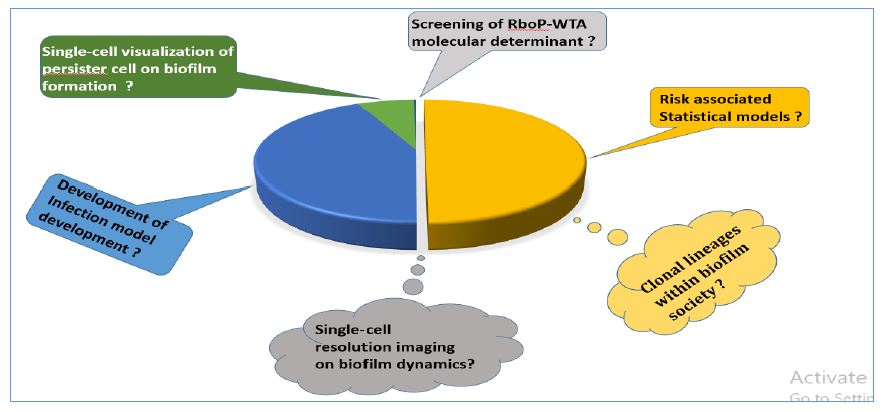
Figures at a glance