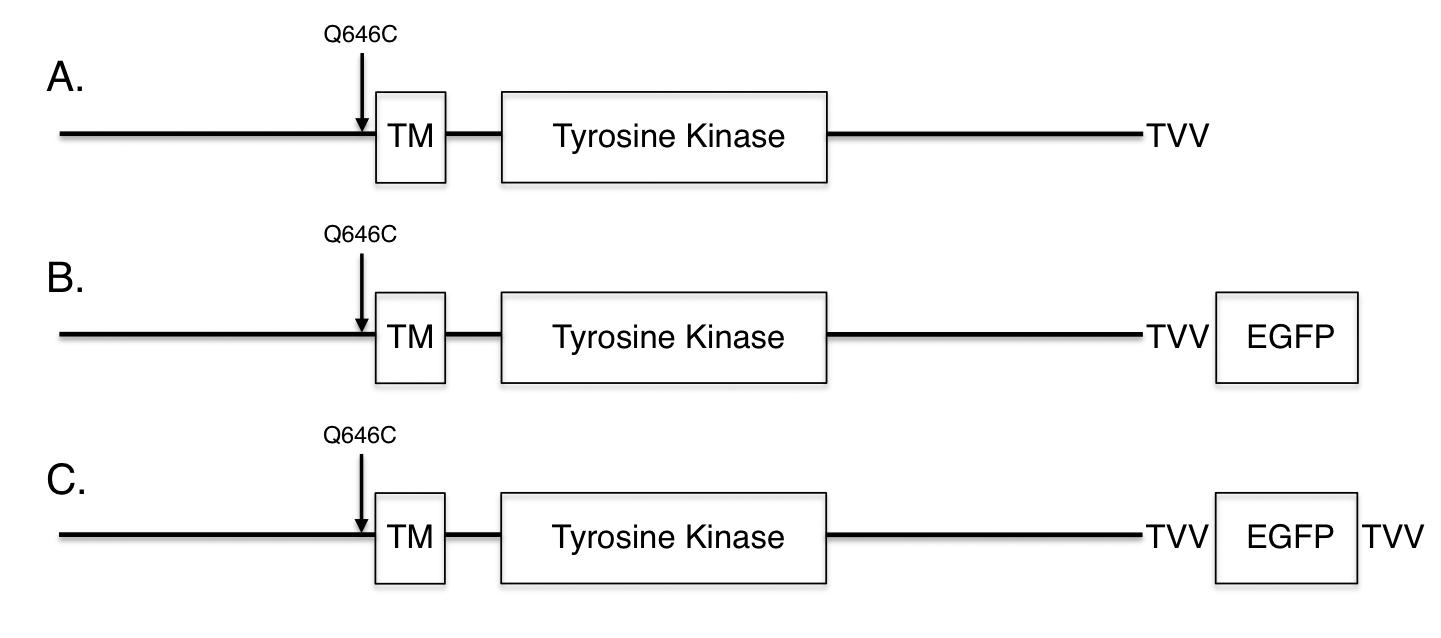
Figure 1: ErbB4 Possesses Multiple Functional Motifs and Mutations Have Been Engineered to Target These Motifs. The organization of ErbB4 is as indicated in this schematic. The extracellular ligand-binding motifs reside in the amino-terminal region upstream of amino acid residue 651. The singlepass transmembrane domain consists of amino acid residues 652-675. The cytoplasmic tyrosine kinase domain consists of amino acid residues 713-989. The majority of cytoplasmic sites of tyrosine phosphorylation reside in amino acid residues 990-1308, most notably Tyr1056. Additional putative functional motifs include a TACE cleavage site, a gamma-secretase cleavage site, two LXXLL (steroid hormone receptor binding) motifs, a BH3 domain, three WW domain binding motifs, and a PDZ domain binding motif. Mutations that disrupt these motifs are noted. Finally, note the two locations of alternative transcriptional splicing, resulting in a total of four different splicing isoforms.

Tables at a glance
Figures at a glance