Construction of the Age-Based DNA Methylation Prognostic Signature for Ovarian Cancer
Received Date: April 11, 2024 Accepted Date: May 11, 2024 Published Date: May 14, 2024
doi: 10.17303/jcrto.2024.12.203
Citation: Li Zhang, Lihua Tang, Fang An, Donghong Zhang, Li Zhou, et al. (2024) Construction of the Age-Based DNA Methylation Prognostic Signature for Ovarian Cancer. J Cancer Res Therap Oncol 12: 1-14
Abstract
Age at diagnosis is a risk for the prognosis of most cancers, whether it works as a prognostic factor in ovarian cancer (OC) remains unclear. Here, we found that age at diagnosis served as a reliable prognostic factor for OC based on the SEER (The Surveillance, Epidemiology, and End Results) database and the survival data from our hospital during 2004–2015. Patients over 65 years old have the lowest overall survival time. According to the coefficients from the Cox regression model, we created age-dependent nomograms for overall survival rates in 3 years, 5 years, and 10 years. The calibration curve and C-statistic values of the nomogram show the optimal consistency between predicted and observed probabilities for all deciles of each age group. Furthermore, the nomogram could effectively distinguish the survival status of high, medium, and low-risk subgroups, which is consistent with best to worst prognosis according to the nomogram risk score. Finally, we constructed an age-based DNA methylation prognostic signature for OC. In conclusion, the developed and verified age-based DNA methylation of nomograms could effectively predict the overall mortality of OC patients of different ages, which will assist clinicians to develop more effective diagnostic strategies and evaluate prognosis.
Keywords: Age; Ovarian Cancer; Prognosis; Nomogram; DNA Methylation
Introduction
Ovarian cancer (OC) is one of the most common gynecological tumors and remains the most lethal cancer of the female reproductive system. According to US statistics, the incidence of OC accounts for 2.5% of all female malignancies, but its deaths account for 5% of all tumors, ranking fifth in female tumor deaths [1]. Whereas in developing countries, the morbidity and mortality of OC rank third [2]. The 5-year survival rate of patients with advanced OC has increased slowly in the past ten years and has not exceeded 40% [3]. Therefore, it is critical to identify the pathological characteristics and prognostic factors of OC [4-7] and improve its survival.
Age at diagnosis is one of the common risk factors for most cancers, including OC, which peaks between the ages of 50 and 80 years. Old women have much higher rates of OC diagnosis and poorer clinical outcomes [8]. Aging is associated with an increased prevalence of frailty, comorbidities, progressive decrease of organ function, as well as adverse drug reactions due to decreasing therapeutic window and distribution volume [9]. Meanwhile, age creates a unique molecular microenvironment for cancer cells, such as metastasis, senescence, inflammation, and apoptosis. However, clinical studies identified that the prognostic effect of age remains controversial [10-12], and the difference in prognostic factors between old and young patients is not evident. For instance, older patients (over 70 years) experienced the same percentage of morbidity with no significant difference in survival when compared with younger (under 70 years) women who were equally debulked [13]. One major reason is they're missing out on the age threshold at the diagnosis of OC [14].
Age-related degeneration of the epigenetic landscape, including progressive loss of DNA methylation over gene-poor genomic regions, which facilitates carcinogenesis [15,16]. Several epidemiological studies have now explored the associations between global hypomethylation and cancer risk, including cancer incidence, mortality and survival [17-19]. Various cancers have been investigated, including lung, breast, colorectal and pancreatic cancers [20]. Evidence suggests that global hypomethylation represents a “mitotic clock” that counts divisions in somatic cells and functions to restrain aging cells and limit malignant progression [21]. Therapies that modulate the pace of methylation loss or eliminate hypomethylated cells could alleviate agingassociated diseases or cancers.
The nomogram has been widely used to estimate the risk probability of death or recurrence for each patient based on the combination of independent prognostic factors to predict a specific endpoint. Up to date, few reports of the nomogram were used to predict the overall survival rate of patients with OC [22-25]. Therefore, our current study aims to reveal the prognostic factors of OC from the perspective of age stratification based on a large number of population studies from the SEER database, and then construct a comprehensive and practical age-dependent nomogram for the prediction of the overall survival of OC patients. Furthermore, we compared the predictive ability of the risk score of different risks for the age-subgroups and provided a reference for predicting the survival status of clinical OC patients. Finally, an age-based DNA methylation prognostic signature for OC was constructed.
Materials and Methods
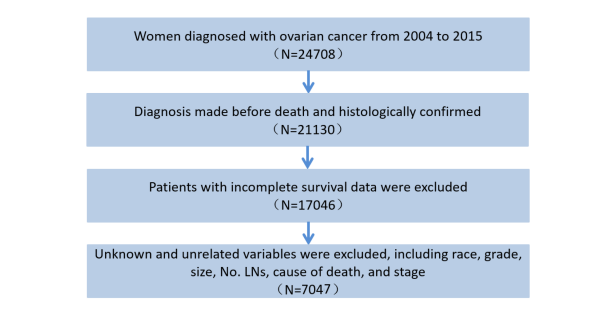
Patient SelectionAll data were collected from the US cancer statistics-SEER database (https://seer.cancer.gov/). We retrospectively analyzed patients with primary OC confirmed by histology. All patients received surgical treatment, and other cases diagnosed by autopsy report or death certificate were excluded. Exclusion criteria include unknown information such as race, tumor size, number of positive lymph nodes (LNs), cause of death, diseases with unknown pathological stage, histology, grade, and incomplete survival data. The process of case selection was shown in Figure 1 and a total of 7047 OC patients were included. We recorded the patient’s age at diagnosis, race, grade, laterality, metastasis, LNs, tumor size, nodes, and AJCC TNM stage for each participant.
We also collected 197 cases of OC patients confirmed by histology during the period from January 1, 2004 to December 31, 2015. We recorded the patient’s age at diagnosis, grade, laterality, the number of positive lymph nodes and 5-year survival values were collected for each participant.
Building a NomogramWe used univariate and multivariate Cox regression analysis to determine whether the age at diagnosis served as independent factors for predicting the OC patients. The Kaplan-Meier method was used to analyze the survival curve and the Log-Rank test was versioned for survival factors statistical analysis [26]. Patients were then divided into three groups according to age. Through the Cox proportional hazards model, variables with significant differences were input into the multivariate analysis for three groups. The independent prognostic factors determined by multivariate analysis were used to construct a nomogram of OS to predict the 3-year, 5-year, and 10-year OS of each group of patients with OC [22,23].
Verification and Calibration of the NomogramTo develop and validate the nomogram model, the patients were randomly divided into training and validation cohorts by applying the ‘createDataPartition’ function in the package of ‘caret’ from R, version 3.6.1. We performed 1000 Bootstrap resampling on the nomogram for internal verification of the training queue and external verification through the verification queue. The C index (concordance index, C-index) was used to evaluate the discriminative performance of the nomogram [27], and measure the prediction accuracy of the model. We carried out the calibration curve to evaluate the degree of calibration of the prediction model, that is, the degree of consistency between the predicted risk of the model and the actual risk.
Survival Rate Analysis Based on Nomogram Risk Group HierarchyThe risk score of the nomogram based on the Cox proportional hazard regression model was calculated[28] and ordered from highest to lowest in three age groups. Three groups of high-, medium- and low-risk were determined and their critical scores were compared by the logrank test.
Construction an Age-Based DNA Methylation Prognostic SignatureThe level 3 DNA methylation (Methylation450k) data and phenotype data of OC were downloaded from UCSC Xena. The β-difference was defined as the difference between mean β value of tumor and normal samples. Then, the student t-test was performed and the p-value was adjusted by Benjamini/Hochberg method. The significant differential methylation level of CpG sites were defined if they met the following criteria of |β-difference| > 0.2 and a false discovery rate (FDR) corrected p-value (Benjamini/Hochberg)< 0.05. The Spearman correlation analysis between age and methylation level of CpG was performed. The Univariate Cox proportional hazards regression model was performed to identify the prognostic value. Furthermore, the least absolute shrinkage and selection operator (LASSO) Cox regression model was performed by R package glmnet (version 4.1-2). The optimal tuning parameter λ was identified via 1- SE (standard error) criterion. According to the risk scores derived from the selected gene signature, a prognosis model was developed.
Statistical AnalysisAll statistical analyses were performed using R software version 3.6.1 and SPSS software version 23.0. The "cmprsk" and "rms" modules in the R software package were used to build models and develop nomograms [29]. Kaplan-Meier curves were utilized to show the impact of each prognostic factor on survival outcomes. P value< 0.05 was considered statistically significant.
Results
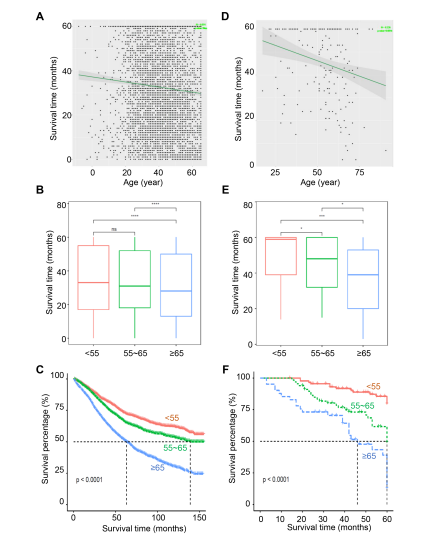
Age at Diagnosis Served as a Reliable Prognostic Factor for OC7047 cases of patients with OC have been included in this cohort study from the SEER database, 2004–2015. To study the role of age at diagnosis on the survival rates of OC, we conducted out the univariate and multiple Cox analysis and found that age was the stable risk prognostic factor for overall OC, when including grade, laterality, histology, lymph nodes, surgery, tumor size, node, metastasis and AJCC (Table 1). In addition, age was negatively associated with five years prognosis of OC (Figure 2A). Then, we cataloged three groups according to the age of OC (Table S1). Significant differences in survival rates were observed among the patients of different ages (p< 0.001, Figure 2B). Patients over 65 years old have the lowest survival time. The Kaplan–Meier curves showed that aging provided effective prognostic factors for the OC patients (p< 0.001, log-rank; Figure 2C). Interestingly, similar results were obtained in OC patients who were collected from Shantou University Cancer hospital in China (Table 2). It is clear that age at diagnosis is a stable risk prognostic factor for OC patients by the univariate and multiple Cox analysis, as well as Kaplan–Meier assay (Figure 2D-F). Our observation indicated that age at diagnosis served as a reliable prognostic factor for OC.
Comparing the Prognostic Impact of Age-DependentTo identify the risk factors for survival rates in different age groups, the univariate and multivariate logistic regression models were derived using the SEER patient data set. As shown in Table S1-2 and Figure S1, the grade, histology, and AJCC were statistically significant for the survival rates of patients less than 55 years old. Likewise, grade, laterality, histology, surgery, LNs, CA125, and AJCC were significant for the middle age patients’ survival. However, for the survival of patients more than 65 years old, all variables were significant except for the laterality, histology, LNs, and node. The race was not found to be statistically significant for all patients.
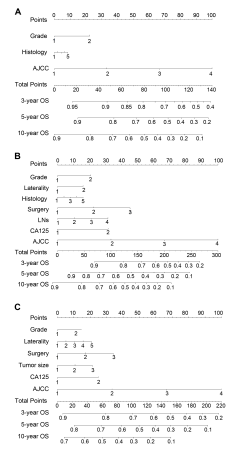
Construction of NomogramThe resulting coefficients from the Cox regression model were then used to create nomograms for the age-dependent overall survival rates in 3 years, 5 years, and 10 years (Figure 3A-C). The factors of interest were considered qualified prognostic factors only if they were significant in both the univariate and multivariate analyses. Interestingly, the higher magnitude odds ratio variables correlated with the largest point allocation on each age nomograms. AJCC was the strongest contributor, whereas grade and histology contributed little to the prognosis of all groups. There were different factors for patients more than 55 years old except for surgery and CA125. For example, laterality and LNs appear the predictors for middle age, but tumor size for old patients, respectively.
Calibration and Verification of NomogramTo calibrate and verify the nomogram, we performed 1000 Bootstrap resampling based on the training queue and external verification. The calibration curve of the nomogram shows that optimal consistency between predicted and observed probabilities for all deciles of each age group in 3, 5, and 10 years, indicating the appreciable reliability of the prognostic nomogram (Figure S3-4). C-statistic values for both training and validation sets were calculated and reported in Table 3. Interestingly, all the values of the C-index were about 0.7. In particular, the C-index values of less than 55 years old were about 0.75 in both the training and testing cohorts. These results demonstrated that the nomograms had excellent discrimination for age-dependent survival of OC patients.
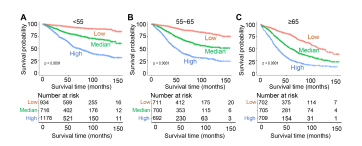
Prediction of the Survival Rate of the Nomogram Risk SubgroupsBased on the Cox proportional hazard regression model, we calculated the total OS nomogram score and divided all patients in each group into three subgroups: lowrisk, intermediate-risk, and high-risk. As expected, the lowrisk subgroup had the best prognosis, while the high-risk subgroup had the worst survival rate in both groups (Figure 4A-C). The results indicated that the nomogram could effectively distinguish the survival status of high, medium, and low-risk subgroups.
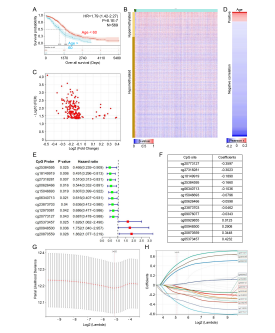
Construction an Age-Based DNA Methylation Prognostic SignatureIncreasing evidence indicates that DNA methylation-based estimates of biologic age can predict important age-related cancer incidence, mortality and survival [20, 30-32]. To construct an age-based DNA methylation prognostic signature in OC, we accessed the DNA methylation sequence data from TCGA database. Firstly, we also found that age at diagnosis served as a reliable prognostic factor for 569 cases of OC, age over 60 patients with a worse survival outcome (Figure 5A). Notably, we identified 1840 CpG sites were significantly changed in OC comparing with normal control group: with 369 hypermethylatied and 1471 hypomethylated CpG sites (cutoff difference>0.2, p< 0.05, Figure 5B). 570 CpG sites were significantly related with the outcomes and 25% of CpG sites were further closely correlated with the age of diagnosis of OC (Figure 5C and D). Moreover, we identified 12 CpG sites cross-linking with the age of diagnosis and prognosis of OC by the univariate Cox proportional hazards regression model (Figure 5E). Furthermore, the least absolute shrinkage and selection operator (LASSO) Cox regression model were performed to construct an age-based DNA methylation prognostic signature (Figure 5F-H). The risk score was developed as a sum of the DNA methylation levels of 13-CpGs sites and their corresponding coefficients, i.e., risk score=-0.359*cg2 0773127+0.423*cg05373457+0.012*cg00929855-0.103*cg06 340713-0.302* cg27318281-0.046*cg23873703-0.034*cg09076077+0.344*cg 20870559-0.188*cg18149919-0.166*cg25384595+0.290*cg0 0948500-0.059*cg00626466-0.079*cg15046693.
Discussion
Based on the poor prognosis of OC, establishing a reliable survival prediction model could effectively guide clinicians to accurately assess the patient's condition and quickly provide targeted treatment plans in a personalized way[25]. In this study, we found that age at diagnosis was an independent risk factor for the prognosis of OC. The age-dependent nomogram has been developed and validated to predict the 3-year, 5-year, and 10-year survival status of patients with OC.
Epidemiological investigations have found that the prognosis of OC may be related to factors such as age at diagnosis, marital status, elevated CA125, tumor differentiation, invasion, lymph node removal, and distant metastasis [33,34]. Consistent with previous studies, tumor grade and lymph node involvement are the comment independent risk factors for ovarian cancer in either group of young or old. While race was a unique independent prognostic factor only in the old group. In addition, we unexpectedly found that the laterality of ovarian cancer was also an independent risk factor for its survival, which conflicts with previous studies [23]. The reason might relate to the criteria of selection and exclusion of cohort studies.
Interestingly, our nomogram model has shown that the AJCC TNM stages were the most contributor to the prognosis for all groups, while the number of lymph nodes is most for the young patients. The reliability of our nomogram was confirmed by the calibration chart, showing the best agreement between the predicted and actual probabilities of its 3-year, 5-year, and 10-year OS. Especially, the C index, as high as 0.732 in the three cohorts showed good discrimination. Furthermore, the subgroups of in subgroups divided by high, medium, and low through prognostic scores, also show the significance of their treatment choices and prognostic judgments. Therefore, the construct nomogram model in our current study is stable and useful for prognostic prediction in all subgroups of OC patients.
To bridge the aging and OC outcome, we constructed an age-based DNA methylation prognostic signature. Consistent with previous studies [16,35,36], our finding indicated a potential mechanistic link between genomic instability, epigenetic age acceleration, and carcinogenesis. Further work is needed to investigate nonlinear trends in epigenetic age that are associated with cancer risk and modeling epigenetic states that are associated with risk of cancer. Integrative analyses of methylation age along with genomic, transcriptomic, and proteomic data within an individual prior to the development of cancer may ultimately be used to develop predictive tools that could be used to guide risk reduction strategies.
Since the SEER database accounts for approximately 28% of the U.S. population, the nomogram we developed has a certain degree of versatility worldwide. The Nomo model we constructed can not only provide a personalized estimate of the prognosis of patients with OC but also could be used by clinicians to make personalized treatment decisions and survival assessments. Unavoidably, our research has some limitations. First, because of lack the of detailed variables in the SEER database, such as race, tumor size, chemotherapy, and cause of death, we excluded about 15,000 cases of OC, and this might result in inevitable inherent biases in our model. Secondly, due to the limitations of the SEER database, some critical survival factors such as FIGO staging, marital status, education, recurrence, radiotherapy could not be studied, which could lower the stability and effectiveness of the nomogram. Third, all include patients were collected from 2004 to 2015, medical changes may affect the results. These strategies include chemotherapy, neoadjuvant chemotherapy, and improvements in targeted therapies. Fourth, it is necessary to validate the nomograms in internal and external cohorts, as well as outside of the SEER program. Therefore, further prospective studies based on other cohorts and multi-center are needed to guarantee the performance of our nomograms.
Conclusions
In summary, we constructed and validated nomograms to predict survival outcomes for age-dependent OC patients, based on a large-scale population from the SEER database. This clinical nomogram could provide a visualized estimation of risk for each prognostic factor and assist clinicians in predicting the 3-, 5-, and 10-year OS for individual OC patients. We also highlight and constructed an age-based DNA methylation prognostic signature for OC.
Supplementary Materials
The following supporting information can be downloaded at: www.mdpi.com/xxx/s1, Figure S1: title; Table S1: title; Video S1: title.
Author Contributions
Conceptualization, Donghong Zhang, Li Zhang, Li Zhou, and Ruimin Li; methodology, Li Zhang, Lihua Zhang and Li Zhou; software, Li Zhang and Fang; validation, Lihua Zhang and Fang An.; resources, Li Zhou, and Ruimin Li; writing—original draft preparation, Donghong Zhang, Li Zhang; writing—review and editing, Donghong Zhang, Li Zhou, and Ruimin Li; supervision, Li Zhou, and Ruimin Li; project administration, Ruimin Li; funding acquisition, Li Zhou, and Ruimin Li. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by grants from the Science and Technology Planning Project of Shantou City, People’s Republic of China (Grant Number: 2019-106). This funding supports the clinical samples collection, data analysis and journal publish.
Institutional Review Board Statement
This study protocol was reviewed and approved by the Cancer Hospital of Shantou University Medical College Ethics Committee, approval number [2021-096]. Written informed consent was obtained from all participants and conformed to the principles in the Declaration of Helsinki.
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study.
Data Availability Statement
All data generated or analyzed during this study are included in this article. Further enquiries can be directed to the corresponding author.
Acknowledgments
In this section, you can acknowledge any support given which is not covered by the author contribution or funding sections. This may include administrative and technical support, or donations in kind (e.g., materials used for experiments).
Conflicts of Interest
The authors declare no conflict of interest.
- Siegel RL, Miller KD, Jemal A (2019) Cancer statistics, 2019. CA Cancer J Clin, 69: 7-34
- Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. (2021) Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin, 71: 209-49.
- Ottevanger PB (2017) Ovarian cancer stem cells more questions than answers. Semin Cancer Biol, 44: 67-71.
- Pei Y, Li K, Lou X, Wu Y, Dong X, Wang W, et al. (2020) miR1299/NOTCH3/TUG1 feedback loop contributes to the malignant proliferation of ovarian cancer. Oncol Rep, 44: 438-48.
- Banerjee S, Moore KN, Colombo N, Scambia G, Kim BG, Oaknin A, et al. (2021) Maintenance olaparib for patients with newly diagnosed advanced ovarian cancer and a BRCA mutation (SOLO1/GOG 3004): 5-year follow-up of a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol.
- Fucikova J, Coosemans A, Orsulic S, Cibula D, Vergote I, Galluzzi L, et al. (2021) Immunological configuration of ovarian carcinoma: features and impact on disease outcome. J Immunother Cancer, 9.
- Moore KN, Chambers SK, Hamilton EP, Chen LM, Oza AM, Ghamande SA, et al. (2021) Adavosertib with Chemotherapy in Patients with Primary Platinum-Resistant Ovarian, Fallopian Tube, or Peritoneal Cancer: an Open-Label, Four-Arm, Phase II Study. Clin Cancer Res.
- Harper EI, Sheedy EF, Stack MS (2018) With Great Age Comes Great Metastatic Ability: Ovarian Cancer and the Appeal of the Aging Peritoneal Microenvironment. Cancers (Basel), 10.
- Tortorella L, Vizzielli G, Fusco D, Cho WC, Bernabei R, Scambia G, et al. (2017) Ovarian Cancer Management in the Oldest Old: Improving Outcomes and Tailoring Treatments. Aging Dis, 8: 677-84.
- Deng F, Xu X, Lv M, Ren B, Wang Y, Guo W, et al. (2017) Age is associated with prognosis in serous ovarian carcinoma. J Ovarian Res, 10: 36.
- Gillen J, Gunderson C, Greenwade M, Rowland M, Ruskin R, Ding K, et al. (2017) Contribution of age to clinical trial enrollment and tolerance with ovarian cancer. Gynecol Oncol, 145: 32-6.
- Gibson SJ, Fleming GF, Temkin SM, Chase DM (2016) The Application and Outcome of Standard of Care Treatment in Elderly Women with Ovarian Cancer: A Literature Review over the Last 10 Years. Front Oncol, 6: 63.
- Wright JD, Herzog TJ, Powell MA (2004) Morbidity of cytoreductive surgery in the elderly. Am J Obstet Gynecol, 190: 1398-400.
- Chan JK, Loizzi V, Magistris A, Lin F, Rutgers J, Osann K, et al. (2004) Differences in prognostic molecular markers between women over and under 45 years of age with advanced ovarian cancer. Clin Cancer Res, 10: 8538-43.
- Minteer CJ, Thrush K, Gonzalez J, Niimi P, Rozenblit M, Rozowsky J, et al. (2023) More than bad luck: Cancer and aging are linked to replication-driven changes to the epigenome. Sci Adv, 9: eadf4163.
- Zabransky DJ, Jaffee EM, Weeraratna AT (2022) Shared genetic and epigenetic changes link aging and cancer. Trends Cell Biol, 32: 338-50.
- Johnstone SE, Gladyshev VN, Aryee MJ, Bernstein BE (2022) Epigenetic clocks, aging, and cancer. Science, 378: 1276-7.
- Li X, Schottker B, Holleczek B, Brenner H (2022) Associations of DNA methylation algorithms of aging and cancer risk: Results from a prospective cohort study. EBioMedicine, 81: 104083.
- Ouchi K, Takahashi S, Okita A, Sakamoto Y, Muto O, Amagai K, et al. (2022) A modified MethyLight assay predicts the clinical outcomes of anti-epidermal growth factor receptor treatment in metastatic colorectal cancer. Cancer Sci, 113: 1057-68.
- Lau CE, Robinson O (2021) DNA methylation age as a biomarker for cancer. Int J Cancer, 148: 2652-63.
- Mangelinck A, Mann C (2021) DNA methylation and histone variants in aging and cancer. Int Rev Cell Mol Biol, 364: 1-110.
- Cham S, Chen L, St Clair CM, Hou JY, Tergas AI, Melamed A, et al. (2019) Development and validation of a risk-- calculator for adverse perioperative outcomes for women with ovarian cancer. Am J Obstet Gynecol, 220: 571 e1- 8.
- Lee CK, Asher R, Friedlander M, Gebski V, Gonzalez-Martin A, Lortholary A, et al. (2019) Development and validation of a prognostic nomogram for overall survival in patients with platinum-resistant ovarian cancer treated with chemotherapy. Eur J Cancer, 117: 99-106.
- Zang RY, Harter P, Chi DS, Sehouli J, Jiang R, Trope CG, et al. (2011) Predictors of survival in patients with recurrent ovarian cancer undergoing secondary cytoreductive surgery based on the pooled analysis of an international collaborative cohort. Br J Cancer, 105: 890-6.
- Rouzier R, Bergzoll C, Brun JL, Dubernard G, Selle F, Uzan S, et al. (2010) The role of lymph node resection in ovarian cancer: analysis of the Surveillance, Epidemiology, and End Results (SEER) database. BJOG, 117: 1451-8.
- Wu L, Zhang D, Zhou L, Pei Y, Zhuang Y, Cui W, et al. (2019) FUN14 domain-containing 1 promotes breast cancer proliferation and migration by activating calcium-NFATC1-BMI1 axis. EBioMedicine, 41: 384-94.
- Chu J, Yang D, Wang L, Xia J (2020) Nomograms predicting survival for all four subtypes of breast cancer: a SEERbased population study. Ann Transl Med, 8: 544.
- Cheng A, Lang J (2020) Survival Analysis of Lymph Node Resection in Ovarian Cancer: A Population-Based Study. Front Oncol, 10: 355.
- Ali AN, Switchenko JM, Kim S, Kowalski J, El-Deiry MW, Beitler JJ (2014) A model and nomogram to predict tumor site origin for squamous cell cancer confined to cervical lymph nodes. Cancer, 120: 3469-76.
- Haghani A, Li CZ, Robeck TR, Zhang J, Lu AT, Ablaeva J, et al. (2023) DNA methylation networks underlying mammalian traits. Science, 381: eabq5693.
- Jung J, McCartney DL, Wagner J, Rosoff DB, Schwandt M, Sun H, et al. (2022) Alcohol use disorder is associated with DNA methylation-based shortening of telomere length and regulated by TESPA1: implications for aging. Mol Psychiatry, 27: 3875-84.
- McCartney DL, Min JL, Richmond RC, Lu AT, Sobczyk MK, Davies G, et al. (2021) Genome-wide association studies identify 137 genetic loci for DNA methylation biomarkers of aging. Genome Biol, 22: 194.
- Lara-Medina F, Perez-Sanchez V, Saavedra-Perez D, Blake-Cerda M, Arce C, Motola-Kuba D, et al. (2011) Triple-negative breast cancer in Hispanic patients: high prevalence, poor prognosis, and association with menopausal status, body mass index, and parity. Cancer, 117: 3658-69.
- Ahmed FE, Ahmed NC, Vos PW, Bonnerup C, Atkins JN, Casey M, et al. (2013) Diagnostic microRNA markers to screen for sporadic human colon cancer in stool: I. Proof of principle. Cancer Genomics Proteomics, 10: 93-113.
- Chen L, Ganz PA, Sehl ME (2022) DNA Methylation, Aging, and Cancer Risk: A Mini-Review. Front Bioinform, 2: 847629.
- Gao T, Joyce BT, Liu L, Zheng Y, Dai Q, Zhang Z, et al. (2016) DNA methylation of oxidative stress genes and cancer risk in the Normative Aging Study. Am J Cancer Res, 6: 553-61.

Tables at a glance
Figures at a glance