PKIB Promotes Breast Cancer Progression Through Immune Infiltration and the GREB1/PKIB Axis
Received Date: January 01, 2025 Accepted Date: February 01, 2025 Published Date: February 04, 2025
doi: 10.17303/jcrto.2025.13.103
Citation: Fengyuan Qian, Sen Zhong, Bowen Zheng, Diya Liu, Yuying Wang, et al. (2025) PKIB Promotes Breast Cancer Progression Through Immune Infiltration and the GREB1/PKIB Axis. J Cancer Res Therap Oncol 13: 1-19
Abstract
Background: PKIB is a critical protein known to function as an inhibitor of protein kinase A. However, its prognostic significance and its relationship with tumor-infiltrating lymphocytes in breast cancer (BC) remain unclear.
Materials and Methods: The mRNA and protein levels of PKIB in breast cancer were assessed. The prognostic value of PKIB was analyzed using data from the TCGA databases. Functional enrichment analysis and immune infiltration analysis were performed. Protein-protein interactions involving PKIB were analyzed using the STRING database. Our in vitro and in vivo experiments confirm the expression and biological functions of PKIB in breast cancer.
Results: PKIB was found to be highly expressed in breast cancer and associated with poor prognosis. The mRNA and protein expression levels of PKIB were correlated with gender, race, age, pathological N stage, ER status, PR status, HER2 status and Menopause status. Functional enrichment analysis revealed that PKIB is involved in extracellular matrix organization, activation of N-methyl-D-aspartate receptors (NMDARs) and postsynaptic events and transmission across chemical synapses. Moreover, PKIB expression was positively correlated with the infiltration levels of macrophages, B cells, Th1 cells and eosinophils in breast cancer. Combining immune infiltration analysis and immunohistochemistry, the GREB1/PKIB axis may be implicated in immune evasion through the regulation of PD-L2 expression. Experimental results indicated that PKIB is upregulated in breast cancer, and inhibiting PKIB can suppress the progression of breast cancer.
Conclusion: The high expression of PKIB in breast cancer is associated with disease progression, poor prognosis, and increased levels of macrophage infiltration. The GREB1/PKIB axis may play a role in the regulation of PD-L2. These findings suggest that targeting the GREB1/PKIB axis or inhibiting PKIB expression may represent a novel therapeutic approach for inhibiting breast cancer progression.
Keywords: PKIB, breast cancer, immune infiltration, GREB1, progression.
Background
Breast cancer (BC) is the most prevalent malignancy in women, with its incidence continuing to rise in recent years [1]. In 2024, it was projected that the United States reported approximately 310,720 new cases of BC, leading to an estimated 42,250 deaths[2]. In China, breast cancer represents about 15.59% of all new cancer cases among women and was one of the top five causes of death among women in 2020 [3]. Characterized by considerable heterogeneity, breast cancer is clinically classified into four subtypes: Luminal A, Luminal B, HER2-positive, and triple-negative [4]. Despite extensive molecular research on breast cancer diagnosis and treatment, the exact molecular mechanisms remain largely unresolved [5,6]. Thus, identifying new therapeutic targets and biomarkers for more accurate diagnosis and prognosis remains a critical focus.
cAMP-dependent protein kinase A (PKA) is widely acknowledged as a key mediator of the extensive physiological and pathological processes triggered by cAMP [7,8]. By interacting with G-proteins, PKA activates various ligand-receptor signaling pathways that govern cellular growth and differentiation [9,10]. The endogenous small peptide inhibitor (PKI) of PKA was found to inhibit the activity of PKA-c kinase and therefore has an important role in carcinogenesis [11]. There are three isoforms of PKI: PKIα, PKIβ (PKIB), and PKIγ. The three isoforms of PKI are small, heat-stable proteins sometimes called PKA inhibitory peptides due to the short sequences that range from 70 to 75 amino acids long [12]. PKIα and PKIγ are both 75 amino acids in length, whereas PKIB is only 70 amino acids long [13-15]. The three PKIs share similar peptide sequences, with the highest similarity being between PKIα and PKIγ, with 96% similarity, whereas PKIα and PKIB share only 40% structural similarity [16,17]. Together, the three isoforms of PKI are known to have nuclear as well as cytosolic functions in oncogenesis and metastasis. All three isoforms of PKI not only have a shared nuclear export sequence but also an affinity to the PKA-c subunits [18]. To date, researches have revealed that PKIB enhances the constitutive activity of the G protein-coupled zinc receptor GPR39 and may play a pivotal role in vascular endothelial cells [19,20].
PKIB has been implicated in several different cancer types as a potential target for cancer therapy. In prostate cancer, the elevation of 17β-estradiol has been implicated in the upregulation of PKIB expression through ERα-mediated nuclear signaling and transcriptional activation [21]. Although the precise mechanisms linking prostate cancer to this pathway remain unclear, PKIB is hypothesized to be a component of the elusive mechanism connecting ERα signaling with the development of castration-resistant prostate cancer. Specifically, the overexpression of PKIB in prostate cancer was found to enhance the invasiveness of cancer cells, whereas the knockdown of PKIB resulted in a marked reduction in cellular invasiveness [21]. Moreover, in osteosarcoma, PKIB has been linked to the expression of two well-known epithelial-mesenchymal transition (EMT) markers, E-cadherin and vimentin [22]. Despite the prevailing evidence that PKIB generally promotes tumor growth, an inverse effect has been observed in colorectal cancer [23]. Additionally, PKIB promotes cell proliferation and the invasion-metastasis cascade through the PI3K/Akt pathway in NSCLC cells [24]. In breast cancer, PKIB expression strongly correlated with phosphorylated Akt expression and also with triple-negative breast cancer subtype [16,25]. Liu et al. postulate that the expression of PKIB may potentiate growth factor receptor signaling pathways and autophagic mechanisms, potentially leading to increased proliferation and drug resistance in breast cancer cells [25]. However, the association between PKIB and immune infiltration in BC has not been fully investigated.
In this study, we investigated the correlation between PKIB expression in breast cancer (BC) and various clinicopathological features, as well as its prognostic significance. Furthermore, we examined the underlying molecular mechanisms and the association between PKIB and tumor-infiltrating immune cells. Collectively, our findings offer novel mechanistic insights into the biological and immunological roles of PKIB, suggesting its potential as a therapeutic target for the prevention and treatment of BC.
Material and Methods
PKIB Expression and Patient Characteristics
RNA-sequencing expression (level 3) profiles and corresponding clinical information for different tumors were downloaded from the TCGA datase (The Cancer Genome Atlas Program database https://portal.gdc.cancer. gov). The data of normal tissues were also obtained from the GTEx database (Genotype-Tissue Expression database https://commonfund.nih.gov/gtex). In addition, PKIB protein expression was analyzed in HPA (The human protein atlas, https://www.proteinatlas.org/) and UALCAN (http://ualcan.path.uab.edu/index.html).
Survival Analysis
The survival package was used for proportional hazards assumption and fitting survival regression, and the results were visualized with the survminer package and ggplot2 package. Best grouping method (best) was selected and the survcutpoint function of survminer package was performed for filtering of the best cut-off [26].
Differential Gene Expression Analysis
According to the median score of PKIB expression, patients with BC in TCGA were divided into high and low expression groups. The DESeq2 package was used to perform the differentially expressed gene (DEG) analysis between the two groups [27]. Adjusted p-value <0.05 and |log2-fold-change (FC) |>1 was set as the thresholds of DEGs. The Ggplot2 package was used to visualize the analysis results.
Functional Enrichment Analysis
After ID conversion of the DEGs, the clusterProfiler package was used for enrichment analysis, including Gene Ontology (GO) analysis, Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis, and Gene set enrichment analysis (GSEA). The Ggplot2 and packages igraph were used to visualize the enrichment analysis results.
Immune Infiltration Analysis
Based on the ssGSEA algorithm provided in R packet-GSVA [1.46.0] [28], the markers of 24 type of immune cells provided in the article [29] were used to calculate the immune infiltration of the corresponding datasets. The analysis results were visualized using ggplot2 package for lollipop map. Spearman’s correlation analysis was performed to investigate the correlation between the expression of the indicated gene and immune cells.
Patients and Samples
A total of 34 BC tissues and matched non-cancerous tissues were collected from breast cancer patients undergoing surgery in Shanghai Tenth People’s Hospital between 2022 and 2024. None of the patients received chemotherapy, radiotherapy, or other neoadjuvant therapy preoperatively. The clinicopathological and molecular characteristics of the patients were described in detail in Table 1. The study received approval from the Ethics Committee of Shanghai Tenth People’s Hospital and was conducted in accordance with Declaration of Helsinki. All patients provided informed consent prior to enrollment. All tissue samples were immediately preserved in liquid nitrogen after removal.
Immunohistochemistry
Immunohistochemistry was carried out as previously described [22] using a PKIB antibody (1:1000, ab233521, Abcam, Cambridge, UK). The intensity of staining was determined as 0 (negative), 1 (weak), 2 (moderate), and 3 (strong).
Cell Culture
Human breast cancer cell lines (BT549, MDA-MB-231, MDA-MB-453, BT474, SK-BR-3, T47D, and MCF-7), human breast normal epithelial cell line (MCF10A) and human embryonic kidney cell line (HEK293T) were obtained from the Chinese Academy of Sciences (Shanghai, China). BT549 and T47D were cultured in RPMI 1640 with 10% Fetal Bovine Serum (FBS, Gibco) and 1% penicillin-streptomycin mixture (Servicebio). MDA-MB-231, SK-BR-3, MDA-MB-453, BT474, MCF-7, and 293T were maintained in DMEM medium with high glucose plus 10% Fetal Bovine Serum (FBS, Gibco) with 1% penicillin-streptomycin mixture (Servicebio). MCF10A cells were cultured in a special medium for MCF10A (Procell). All cell lines described above were cultivated at 37 °C with 5% CO2.
Cell Transfection and Stable Cell Line Establishment
The short hairpin RNAs (shRNAs) plasmids were designed and synthesized by GeneChem (Shanghai, China). The LentiFit transfection kit (GeneChem, Shanghai, China) was applied to establish stable cell lines and selected using 2 µg/ml puromycin. The transfection of plasmids was performed using Lipofectamine 3000 (Invitrogen), according to the manufacturer’s instructions.
CCK8 and Colony Formation Assay
The ability of cell proliferation was evaluated by CCK8 assay. Transfected BC cells (BT549 1.5 ×103 cells/well; MDA-MB-231 2.0×103 cells/well) were seeded into 96-well plates. CCK8 reagent (10 μg) (Yeasen, China) was added into each well and cells were incubated for 2h at 37°C in 5% CO2. Then absorbance at 450nm was recorded. For colony formation assay, transfected cells (BT549 0.8 × 103 cells/well; MDA-MB-231 1.0 ×103 cells/ well) were seeded into 6-well plates. The medium was replaced every 3–4 days. After culturing 7 to 14 days, colonies were fixed in 4% paraformaldehyde and stained with 0.01% crystal violet.
Cell Migration and Invasion Assay
For migration assay, the treated cells (BT549 4×104 cells; MDA-MB-231 5×104 cells) were seeded into the upper chamber of Transwell® permeable supports (Corning Costar Corp, Cambridge, USA), whereas, for invasion assay, BC cells (BT549 8 ×104 cells; MDA-MB-231 1.0×105 cells) were placed in the upper chamber with matrigel (BD Biosciences, USA). After incubation for 18 h, non-migrated or invaded cells were gently wiped off with a cotton swab and cells in the bottom chamber were fixed with 4% paraformaldehyde, and stained with 0.01% crystal violet. Then five visions were photographed by a microscope at a magnification of ×200 and counted with ImageJ software. The experiments were replicated in triple.
Xenograft Subcutaneous Implantation
Four-week-old BALB/c nu/nu female mice were purchased from Shanghai Lingchang Biotechnology (Shanghai, China). A total of 1 × 107 MDA-MB-231 cells after stable transfected with shPKIB or Ctrl were subcutaneously injected into each mouse. The mice were euthanized on the 14th days after injection. Then the tumor weight and volume were measured. All animal studies were approved by the Institutional Animal Care and Use Committee of the Shanghai Tenth People’s hospital.
Statistical Analysis
All statistical analyses were conducted using GraphPad Prism 8.0 (GraphPad Software, San Diego, CA, USA) or R version 4.2.1 (R Core Team, http://www.r-project.org/). Wilcoxon rank sum test and Wilcoxon signed rank test were used to analyze the statistical significance for the PKIB expression in the non-paired and paired samples, respectively. Survival analyses were conducted using the Kaplan–Meier method. Other statistical analyses of data of different groups were performed using the Student’s T test. A pvalue <0.05 was considered to indicate statistical significance.
Results
PKIB is Increased in BC Tissues
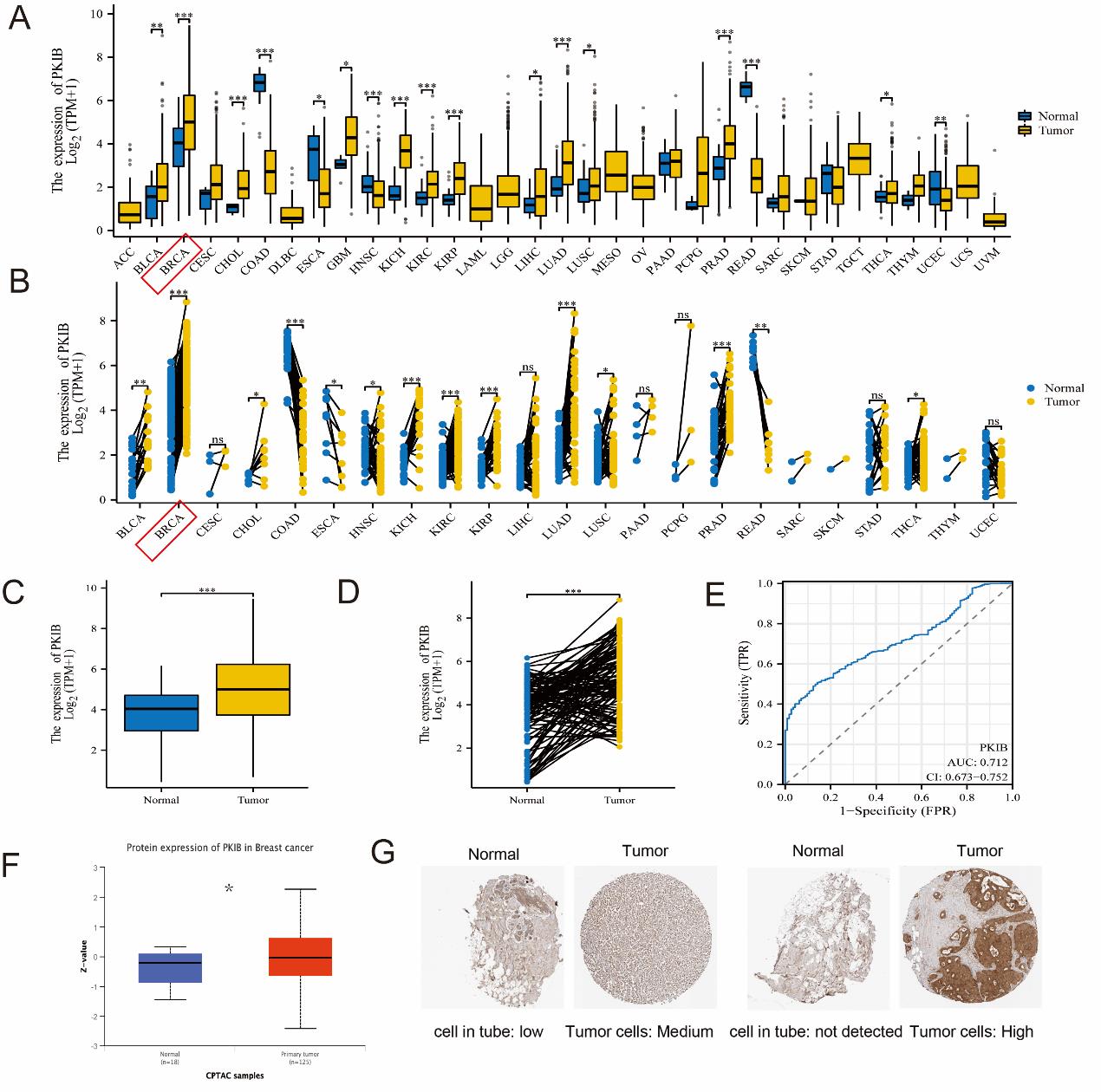
To explore differences of PKIB expression between tumor and normal tissues, we performed the pan-cancer analysis, which showed that PKIB was highly expressed in most tumors in TCGA compared with unpaired normal tissues respectively, such as breast cancer (BRCA), lung adenocarcinoma (LUAD), kidney renal clear cell carcinoma (KIRC), and prostate cancer (PRAD) (Figure 1A). We further compared PKIB expression between paired tissues in TCGA. Interestingly, PKIB was up-regulated in BRCA, bladder urothelial carcinoma (BLCA), KIRC, kidney papillary cell carcinoma (KIRP), LUAD and PRAD. However, PKIB expression was significantly lower in rectum adenocarcinoma (READ), esophageal squamous cell carcinoma (ESCA) and colon adenocarcinoma (COAD) (Figure 1B). Additionally, PKIB is highly expressed in BC tissues compared not only with unpaired normal tissues but also with paired normal tissues (Figure 1C and D). Furthermore, receiver operating characteristics (ROC) analysis indicated that PKIB might serve as a biomarker for distinguishing BC from normal tissues with an area under the ROC curve (AUC) of 0.712 (95%CI=0.673–0.752) (Figure 1E). Besides mRNA expression, PKIB protein is increased in BC tissues compared with normal tissues (Figure 1F). Immunohistochemical (IHC) staining also showed that PKIB protein expression is up-regulated in tumor cells compared with cells in normal tissues from the HPA database (Figure 1G). Collectively, these results indicated that PKIB is increased in BC and might be a novel diagnostic biomarker.
PKIB mRNA Expression is Associated with Clinic-pathological Characteristics
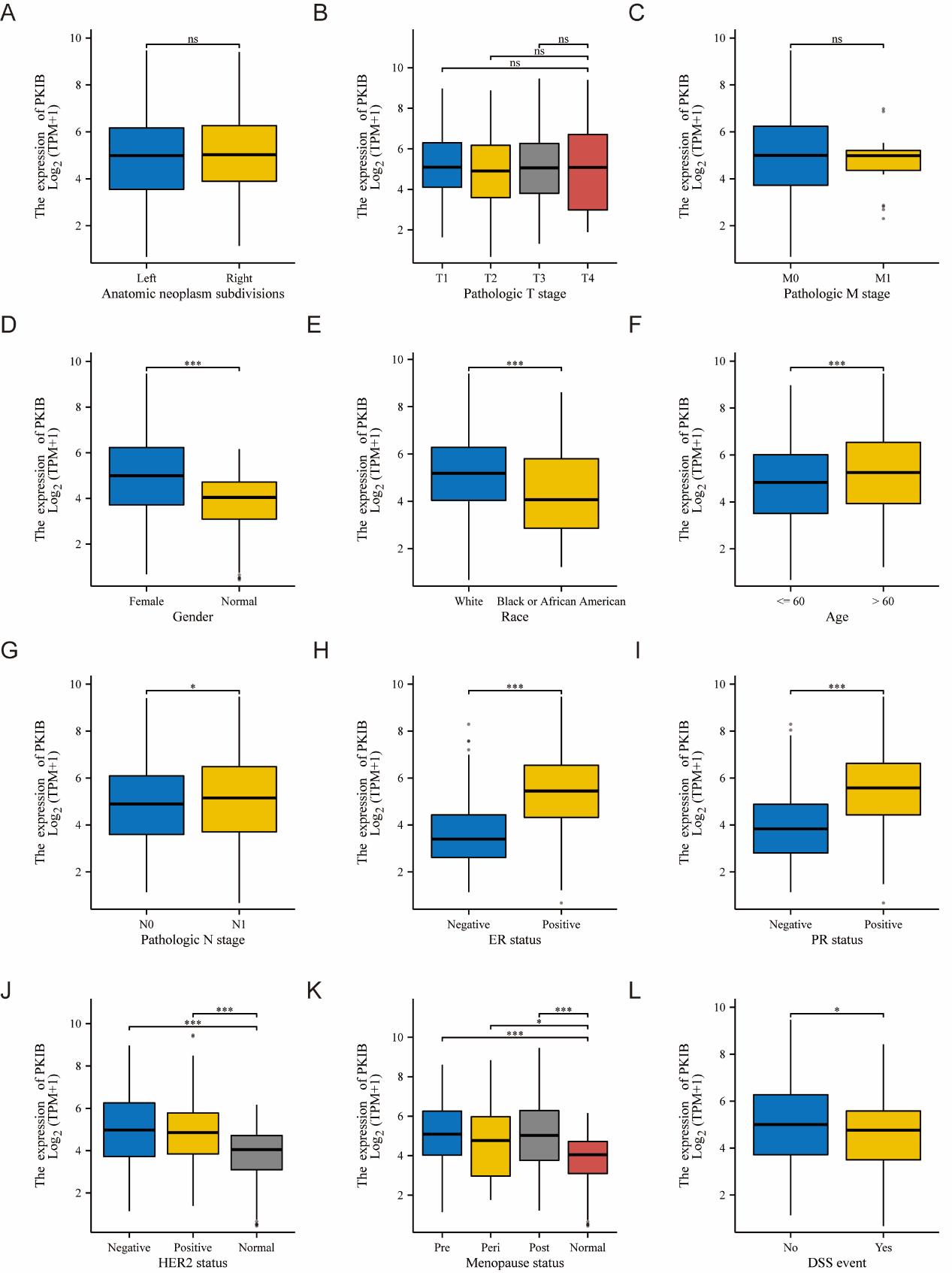
High expression of PKIB was significantly correlated to disease-specific survival (DSS) events (p=0.019), overall survival (OS) (p=0.051), and progress-free interval (PFI) events (p=0.048) (Table S1). The mRNA expression of PKIB in different subgroups of BC was further evaluated, and the results suggested that PKIB expression was not corrected with patient anatomic neoplasm subdivisions, pathological T stage, and pathologic M stage (Figure 2A-C), whereas PKIB expression was correlated with gender, ethnicity, age, pathological N-stage, ER-status, PR-status, HER2-status, menopausal-status and DSS events (Figure 2D- L).
PKIB Protein Level is Associated with Clinicopathological Characteristics
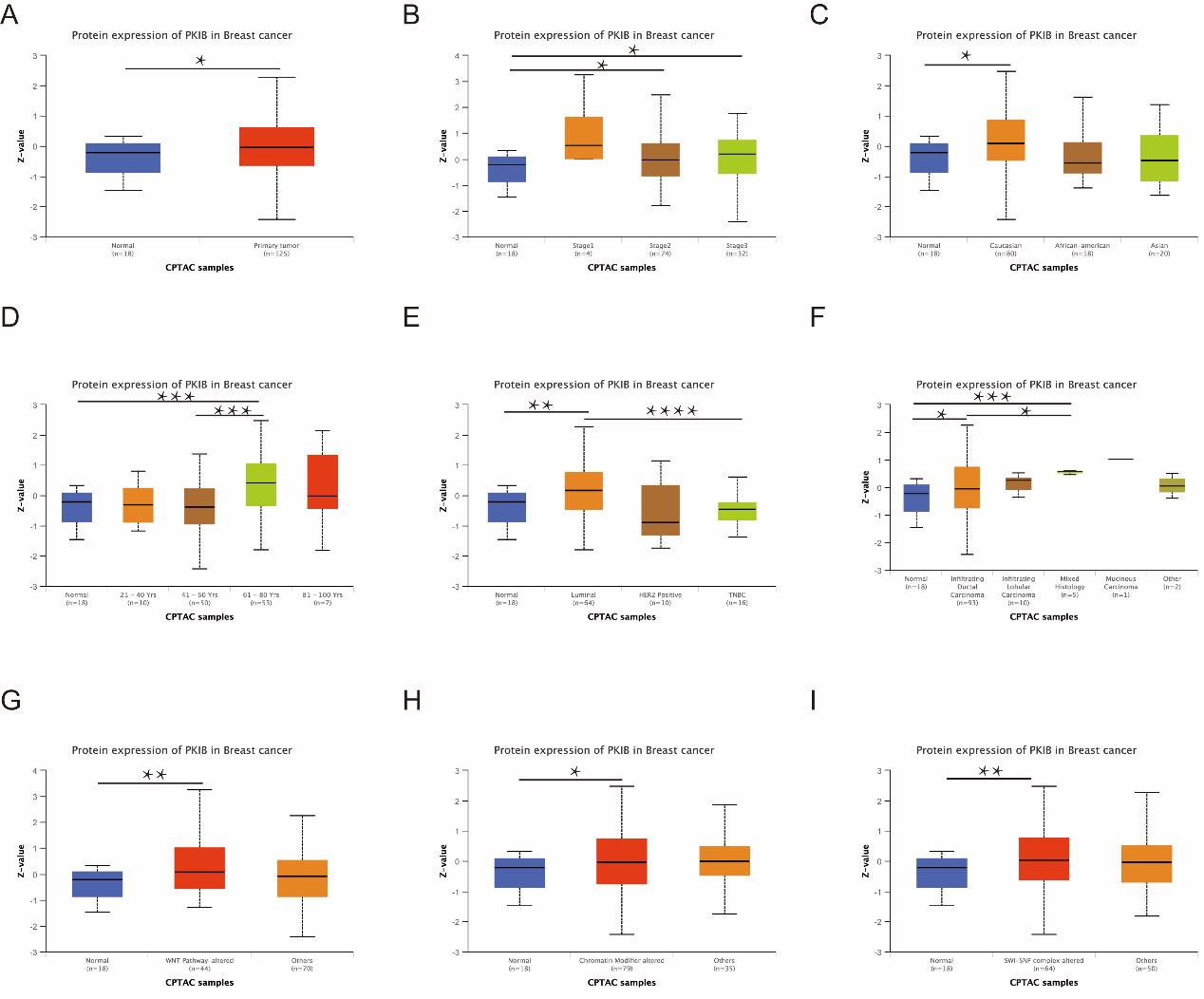
In addition to mRNA expression, protein expression in different subgroups of BC was determined by the ULCAN database. We analyzed PKIB protein exposure in breast cancer using CPTAC samples and compared it with that of normal tissue (Figure 3A). The outcomes demonstrated that the high level of PKIB expression was not associated with the cancer stage and race of the patients (Figure 3B-C), whereas PKIB expression was significantly correlated with age, major subclasses, and tumor histology (Figure 3D and E). In addition, we found that the expansion of PKIB expression was significantly over-regulated in samples with altered WNT pathway, altered chromatin modifiers, and altered SWI-SNF complexes (Figure 3F-H).
Low PKIB Expression was Negatively Correlated with Poor Prognosis
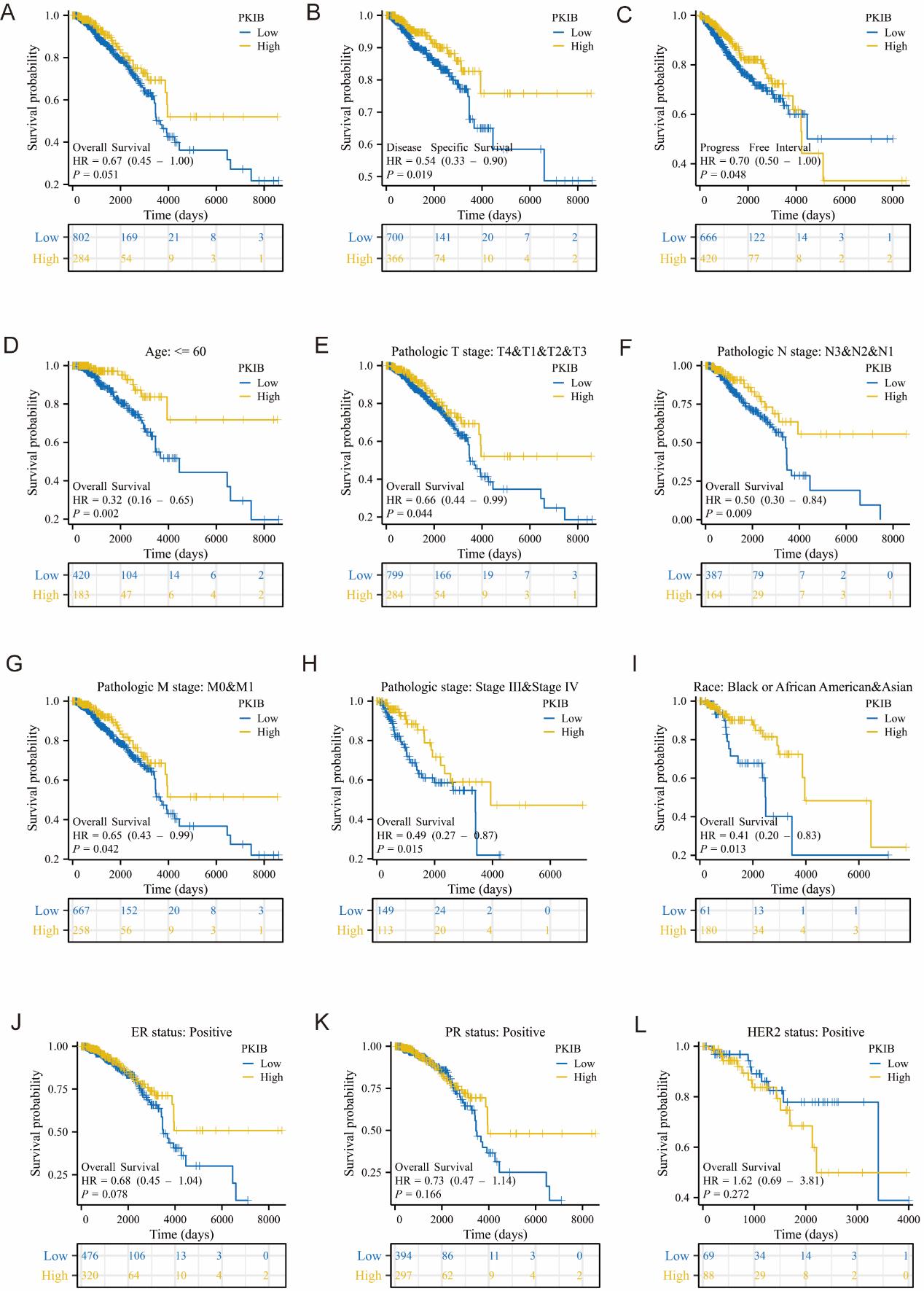
To evaluate the impact of PKIB (PKIB) observation on patients' prognosis, we performed survival analysis of BC patients. BC patients. Interestingly, the outcome indicated that low protein expression of PKIB was accompanied by poorer OS, DSS, and PFI (Figure 4A-C). Similar to these results, low expression of PKIB was related to a worse OS in different subgroups, including in patients of less than 60 years old, T3 and T4 stages, N3 and N2 and N1 stages, M0 and M1 stage, stage III and IV, Asian and Black or African American, (Figure 4D-I). However, PKIB expressed was not significantly correlated with OS in patients with ER positive and PR positive (Figure 4J-L). Consequently, the low PKIB expression of PKIB was relevant to poor OS in basal affected patients (Figure S1D). Moreover, we analyzed the BC dataset of German Cancer Research Center (DKFZ) in PrognoScan. The survivorship analysis (Kaplan–Meier) showed a positive association between less expression of PKIB and poor OS of BC patients (Figure S1E).
Functional Enrichment Analysis Of PKIB-Related Differentially Expressed Genes (DEGs)
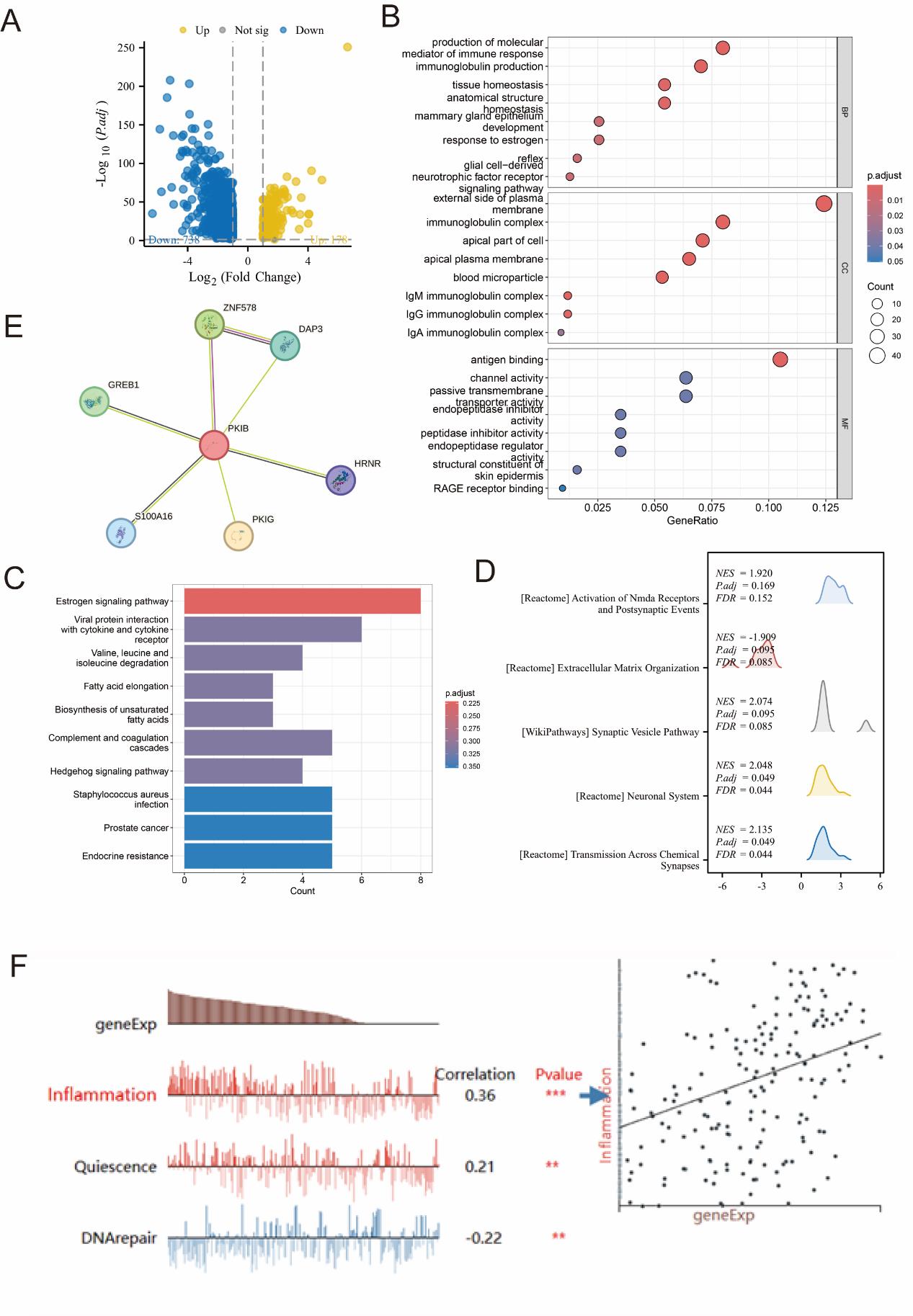
To further explore the underlying mechanism of the PKIB gene in BC, we first identified the PKIB-related differentially expressed genes, comprising 178 up-expressed genes and 738 down- expressed genes (Figure 5A). Next, GO and KEGG analyses suggested that DEGs were enriched in distinct GO aspects, like molecular mediator of immune response generation, external side of plasma membrane outer wall, and in conjunction with binding (Figure 5B). In addition, KEGG pathways showed that DEGs were enriched in estrogen signaling pathway, in the complement and coagulation cascades, and in the interaction of viral proteins with cytokines and cyclin receptors (Figure 5C). Subsequently, GSEA indicated that the PKIB high-expression group was enriched in extracellular matrix organization, NMDA receptor activation and expression of presynaptic events, chemical synaptic transmission, and synaptic vesicle pathways (Figure 5D). Finally, protein-protein interaction network of PKIB and other related proteins were analyzed in the STRING database and the results suggested that PKIB could interact with ZNF578, DAP3, GREB1, HRNR, PKIG and S100A16 (Figure 5D). Previous studies reported that PKIB and GREB1 interactions can mediate BC cell estrogen response, promote breast cancer cell proliferation and drug resistance, and promote PI3K/Akt signaling pathway and autophagy [25,30]. Tumor heterogeneity can be better estimated using single-cell sequencing technology [31]. Based on the CancerSEA database [32], a study of BC single cells found PKIB expressed positive correlation with inflammation, DNA repair, and quiescence in BC (Figure 5E).
PKIB Expression Correlated with the Level of BC Immune Infiltration
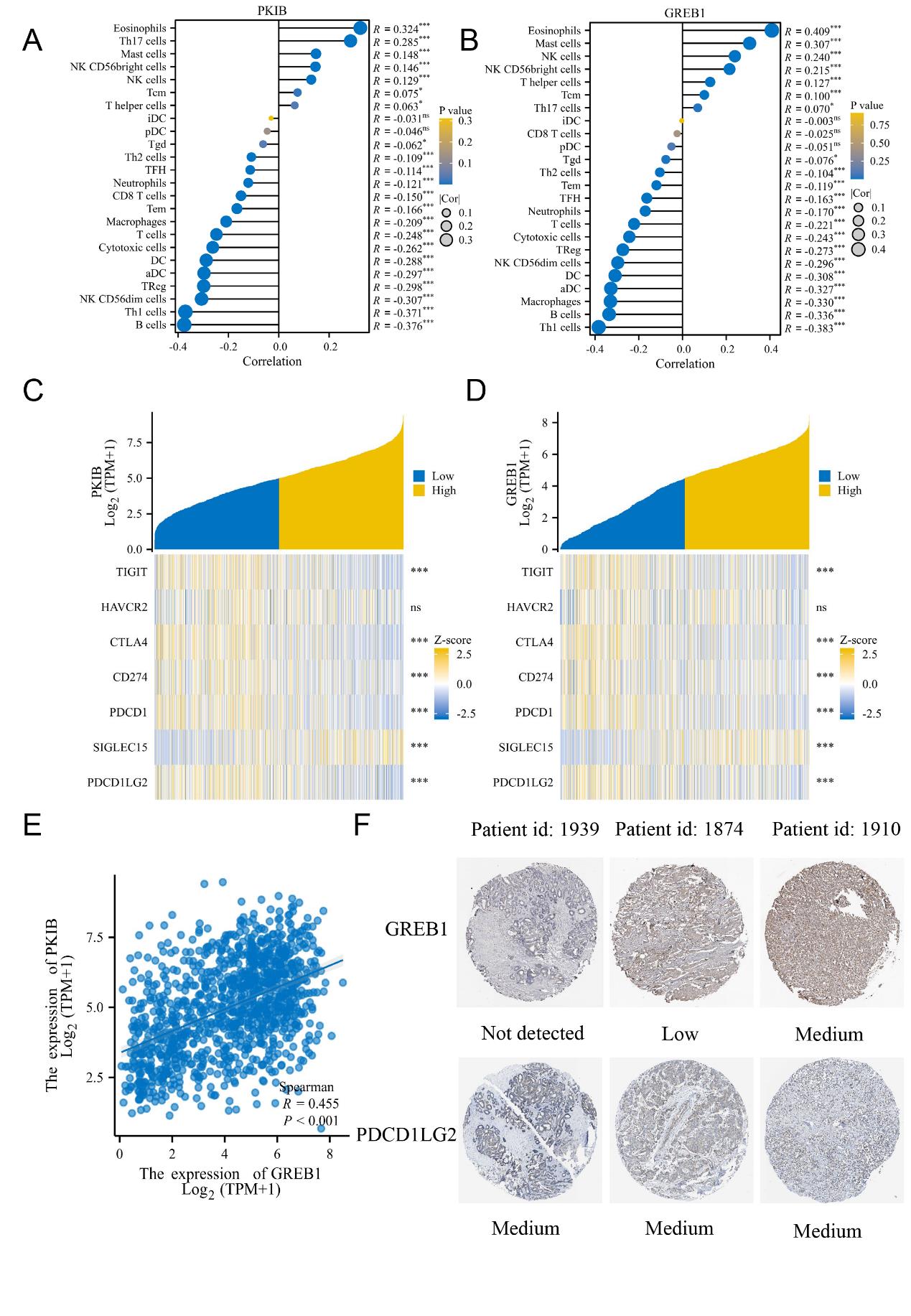
Tumor microenvironment has a significant number of immune cells. Consequently, we investigated if the PKIB expression correlates with the infiltration of immune cells. The results indicated a significant negative correlation between PKIB expression and the level of macrophage infiltration (r=-0.209, p<0.001), B cells (r=-0.376, p<0.001), and Th1 cells (r=-0.371, p<0.001). Nevertheless, there was a positive correlation with the level of eosinophilic infiltration (r=0.324, p<0.001) (Figure 6A). Previous data has demonstrated that GREB1 is an evolved conserved protein whose glycosyltransferase structural domain links ER α glycosylation to cancer stability (19, 20). Considering our protein-protein interaction network result (Figure 5D), we examined the relevance of GREB1 to immune cells in BC. As illustrated in Figure 6B, GREB1 expressed was also significantly negatively related to the levels of macrophages infiltration (r=0.330, p<0.001), whereas it was positively related to the infiltrating levels of Eosinophils (r=-0.409, p<0.001) (Figure 6B, Figure S3A, and S3B).
To reveal the correlation between PKIB or GREB1 expression and immune-checkpoint-related gene expression, heatmaps of the correlation were generated and are shown in Figure 6C and D. The results suggested that PKIB expression was positively corrected with TIGIT, HAVCR2, CTLA-4, CD274 (PD-L1), PDCD1 (PD-1), SIGLEC15 and PDCD1LG2 (PD-L2) (Figure 6C). However, GREB1 expression was positively correlated with PDCD1 and PDCD1LG2 (Figure 6D). Since PKIB expression was positively correlated with GREB1 expression, we focused on their common immune-checkpoint-related gene, PDCD1LG2 (Figure 6E, Figure S3C, and S3D). As an evolutionarily conserved protein, GREB1 could bind to PKIB to modify signaling [33]. Interestingly, we observed the positive correlation of protein expression of GREB1 and PDCD1LG2 in BC tissues from the HPA database (Figure 6F). Taken together, our results suggested that PKIB acts a site-specific role in immune infiltration in BC and the PKIB/GREB1 axis might be a regulator of PDCD1LG2.
PKIB Protein Expression in BC Tissues and its Associations with Clinicopathological Parameters in our Hospital
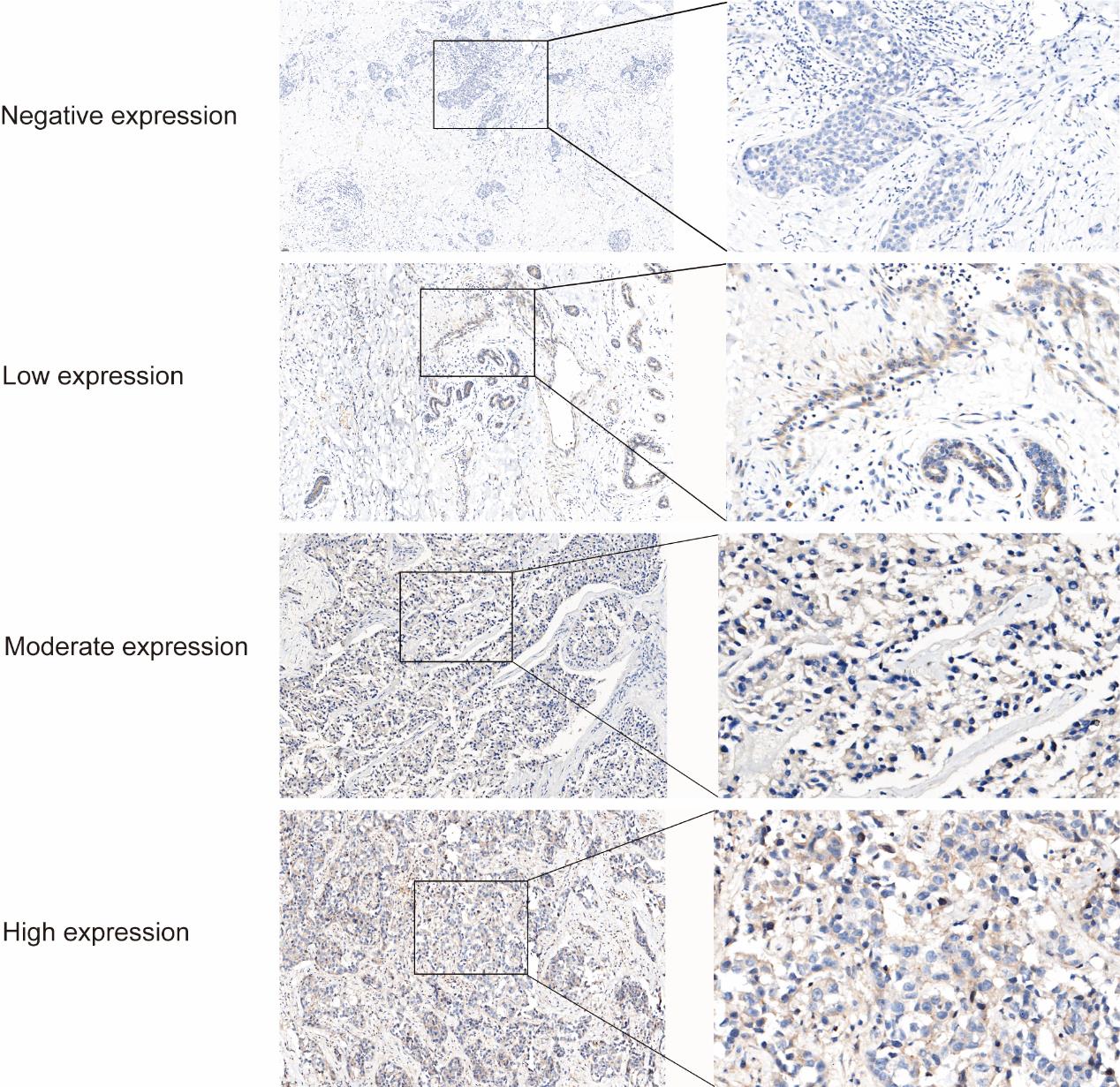
The clinical-pathological features of the cohort are summarized (Table 1). IHC staining was used to examine the expression level of PKIB protein in the BC tissues of 34 cases. PKIB protein staining showed that PKIB was overexpressed in 40% (16/40) of the BC tissues (Figure 7). The subsequent analysis of the association between PKIB expressions and clinicopathological parameters showed that high PKIB expression correlated with any other clinical-pathological values (Table 1).
PKIB Downregulation Suppresses BC Cell Growth, Migration, and Invasion
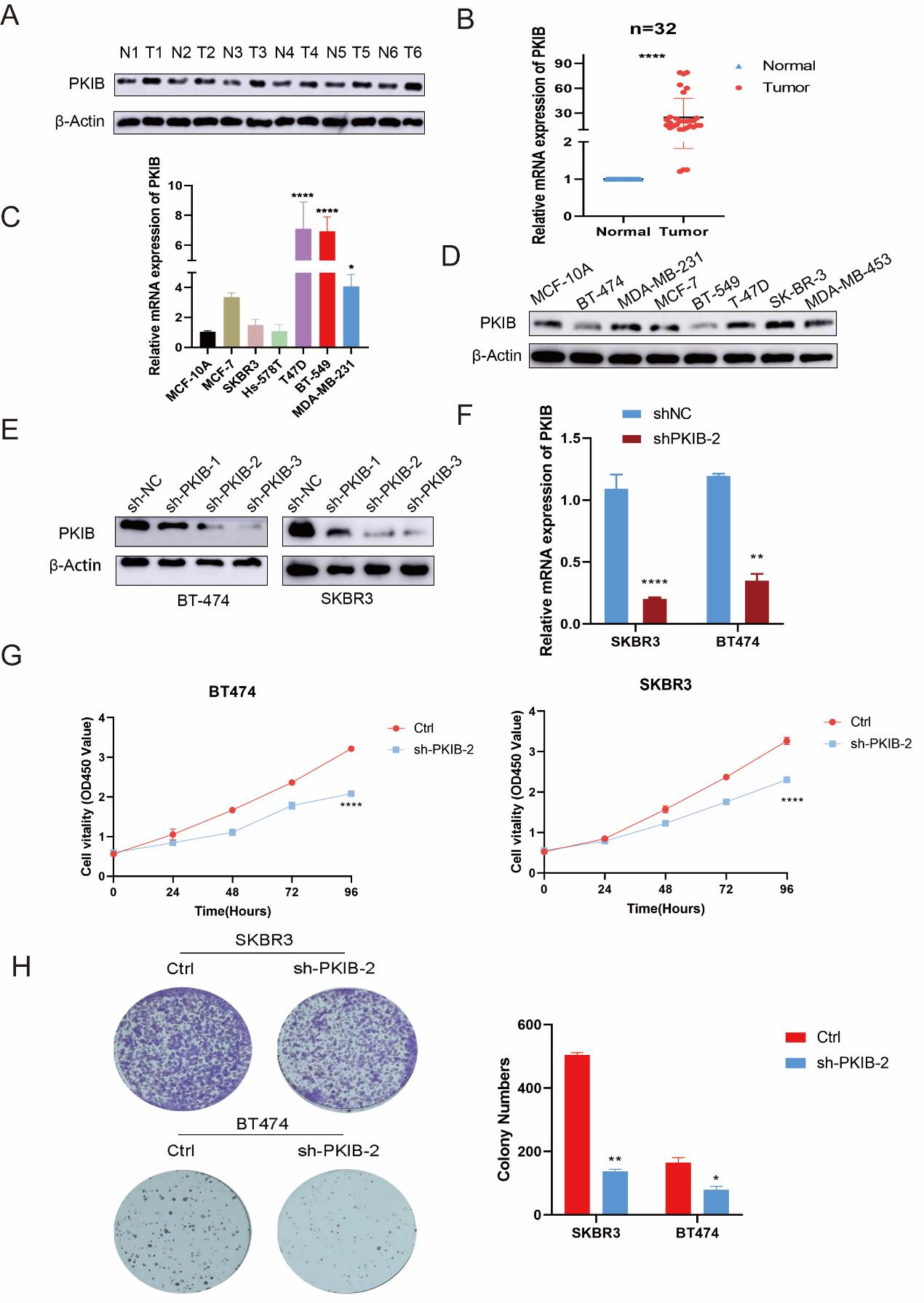
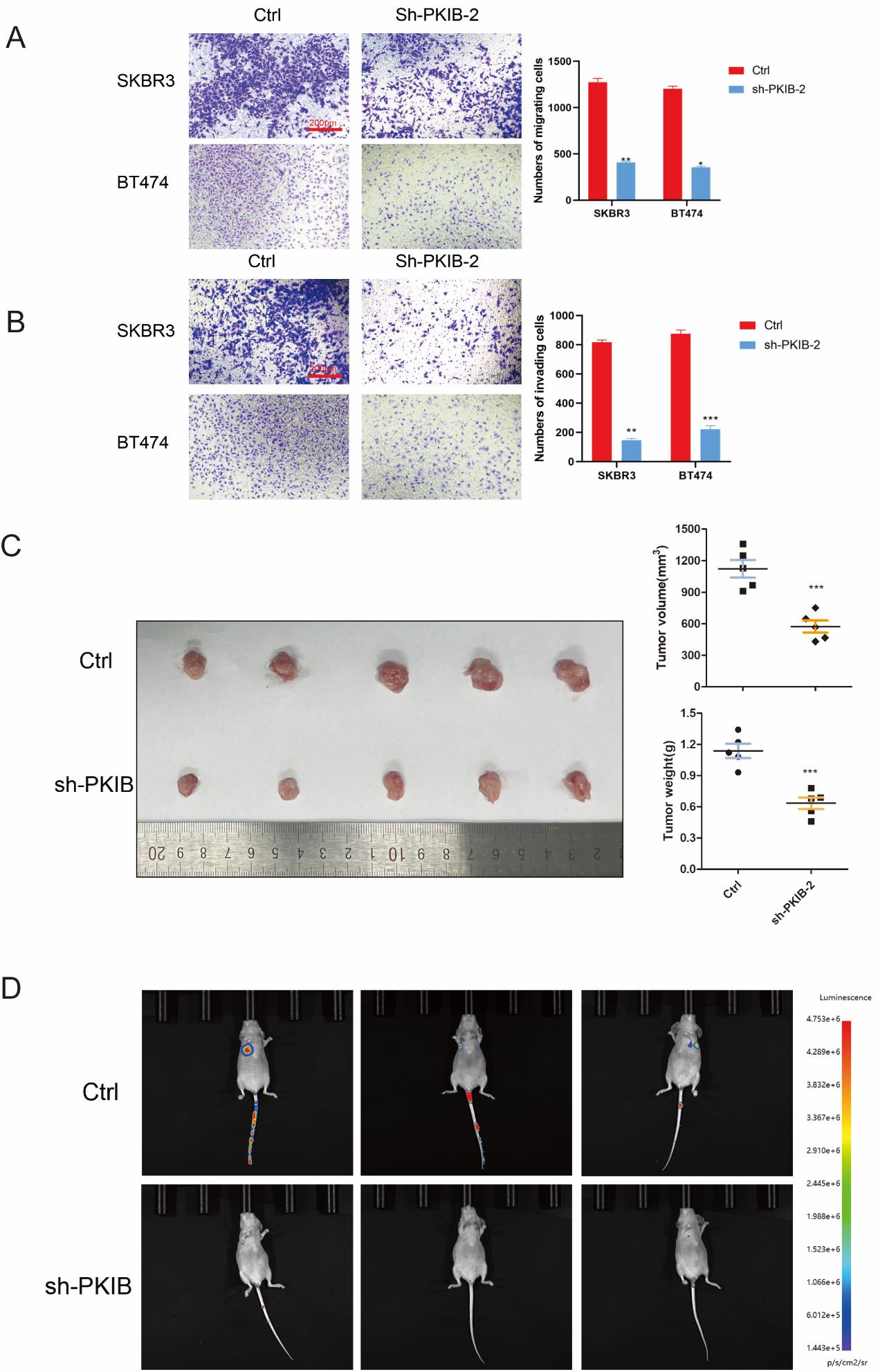
Firstly, western blot (WB) analysis revealed that PKIB levels were dramatically raised in tumors compared to the normal tissues (Figure 8A). Furthermore, the expression of PKIB in BC cells was assessed against that in normal breast epithelial cells using qRT-PCR and WB assays. The results indicated that PKIB expression is markedly increased in BC cells relative to MCF-10A cells (Figures 8B, C). To elucidate the biological function of PKIB in BC cells, we established stable knockdown cell lines for PKIB. Initially, the expression of PKIB in these stably transfected cells was confirmed through qRT-PCR and WB assays (Figures 8D, E). A range of in vivo and in vitro trials were conducted to investigate the biological role played by PKIB within BC cellular contexts. Downregulation of PKIB dramatically inhibited the proliferation of BT549 and MDA-MB-231 cells as verified using Cell Counting Kit-8 (CCK-8) (Figure 8F). Additionally, the colony formation assay was performed to assess the prolonged effects of reduced PKIB on BC proliferation. The number of colonies was consistently lower in the PKIB down-regulated group compared to the vector control group (Figure 8G). In addition, we have characterized the impact of PKIB on BC cell migration and invasion. Our observations revealed that downregulation of PKIB significantly inhibited the migratory and invasive capabilities of both BT549 and MDA-MB-231 cells compared to controls (Figure 9A, B). The results of these in vitro studies suggest that PKIB functions as an oncogenic driver in BC cell lines. To explore further the role of PKIB in tumor development in vivo, we subcutaneously injected MDA-MB-231 cells with stably reduced expression of PKIB or vector-transfected cells into nude mice for tumor model studies. As anticipated, tumor growth was apparently inhibited in the PKIB downregulation group relative to the vehicle group. Furthermore, at the conclusion of our observation period, both tumor size and mass were dramatically diminished in the PKIB downregulation group as compared to the respective controls (Figure 9C). Additionally, a lung metastasis model indicated fewer lesions present in the sh-PKIB groups (Figure 9D). In summary, these combined in vitro as well as in vivo data demonstrate PKIB act as a critical oncogene by facilitating tumor promotion and playing an essential role in BC metastasis.
Discussion
Due to the critical importance of targeted therapy for breast cancer (BC), urgent need to the develop of new early biomarkers and therapeutic targets [34]. PKIB has attracted extensive attention in recent years due to its abnormal expression and function in various cancers, as well as an inhibitor of PKA [35]. Recent studies have shown that PKIB is increased in prostate cancer, non-small cell lung cancer, breast cancer and osteosarcoma [21,22,24,25]. Although previous reports have documented high expression levels of PKIB in BC [16,25], a comprehensive analyse of the relation between PKIB and clinical feature of BC individuals remains lacking. This study aims to elucidate the prognostic value of PKIB expression, explore its relationship with immune infiltrates, and investigate potential molecular mechanisms underlying PKIB's role in BC.
In this investigation, we primarily utilized online databases alongside our own patient data due to limitations posed by small sample sizes and a scarcity of clinical information. Analysis of the PKIB expression in BC using several online databases revealed that PKIB had higher mRNA and protein levels in BC tumors tissues compared to normal tissues. Furthermore, analysis revealed that high PKIB mRNA expression was associated with gender, race, age, pathological N stage and DSS of BC patient, in agreement with the previous study [25]. However, we found PKIB expression was not associated with pathological M stage, which is contrary to previous conclusion [16]. This discrepant result is likely due to differences in study designs and sample sizes. In addition, our results indicated that PKIB is not only a predictive marker, but also a prognostic marker. This has been reported in many other cancers, which include cancers of the prostate, non-small cell lung cancer, bone sarcoma, and breast cancer [21,22,24,25]. PKIB as a key cell-extracellular matrix mediator (ECM), has been shown to enhance tumor regression and metastasis [22]. Nevertheless, there are rare studies on the mechanism of action of PKIB in BC. Our data revealed that PKIB was a regulator of extracellular matrix and multiple signaling pathways, including EMT, inflammation, DNA repair, and quiescence in BC. These fundings provide insights into the mechanism of BC development and deserve further study.
BC has been reported to be a highly immune infiltrated tumor [36]. Given that immunotherapy has shown remarkable clinical efficacy in BC [37] and PKIB is also expressed on a variety of immune cells. Therefore, it is essential to investigate the relationship between immunological infiltration and PKIB expression in BC. TAMs are known to facilitate tumor growth and regulate tumor immunity by producing cytokines [38]. Additionally, advanced BC is associated with increased infiltration of helper T cells 2 (Th2) and T regulatory cells (Tregs) , which may lead to the immune escape of tumor cells [39]. In the present study, we found that PKIB expression was significantly correlated in a positive way with the level of eosinophil infiltration and in a negative way with the level of macrophage infiltration, B cells, Th1 cells. These results suggested that PKIB may indirectly affect the immune infiltration of BC.
Immune checkpoints are specific targets for immunotherapies. Therefore, we also analyzed the correlation between PKIB and immune checkpoints and found PKIB is positively correlated with several immune checkpoints, including PD-L2. Single-cell RNA sequencing (scRNA-seq) indicated that the GREB1/PKIB interaction was involved in a complex cross talk between malignant cell clusters, TAMs, and CD8+T cells, while the PD-1/PD-L2 interaction was identified between CD8+T cells and TAMs. Meanwhile, PD-L2 is upregulated in macrophages and tumor cells after immune checkpoint blockade (ICB) [40]. PD-L1 and PD-L2 are the two types of ligands for PD-1 and the interplay between PD-1 and PD-L1/PD-L2 contributed to the inhibition of T-cell function [41]. Positive correlations between PKIB, GREB1, and PD-L2 were found in BC in our study. Thus, we hypothesized that the interaction between PKIB and GREB1 is involved in immune escape by regulating PD-L2 expression, which suggested blocking GREB1/PKIB axis may be important for immunotherapy in BC.
A great deal of evidence implicates the function of PKIB in tumor progression [35]. Our data demonstrated that PKIB is positively correlates with BC progression, and the knockdown of PKIB expression suppresses the progression of BC cells. Such findings indicate that PKIB may be a potential therapeutic target for BC. While this study has significantly expanded our understanding of PKIB's role in BC, several limitations must be considered. Notably, the study primarily utilized bulk data, which may have obscured interindividual heterogeneity. Consequently, our results require additional validation in a large-scale, real-world cohort. Furthermore, more experimental evidence is needed to elucidate the precise function and mechanism of PKIB in BC. Overall, our study highlights the prognostic value and functional role of PKIB in BC, especially its association with the infiltration of immune cells. Additionally, we identified that the GREB1/PKIB axis may participate in the immune response by regulating PD-L2 expression. These findings suggest that PKIB is a biomarker of prognosis in relation to BC immune infiltration and metastasis. However, further research is necessary to fully elucidate the mechanism of action of PKIB.
Conclusion
In summary, our study indicates that overexpression of the PKIB protein occurs in BC and suggests it could be treated as an independent prognostic biomarker for this malignancy. The knockdown of PKIB expression exhibited a multifaceted inhibitory effect on breast cancer cell progression via the extracellular matrix-receptor interaction pathway. Overall, our findings may provide valuable insights into potential biomarkers for diagnosis and targets for treatment of breast cancer.
Funding
This work was supported by the National Natural Science Foundation of China (no.82073204), the Shanghai Municipal Health Commission (no.202040157).
Ethics Statement
It is approved by the Ethics Committee of Shanghai Tenth People’s Hospital, Shanghai, China.
Data Availability Statement
The original data presented in the study are included in the article, further inquiries can be directed to the corresponding author.
Author Statement
Fengyuan Qian: Writing–review & editing, Writing–original draft, Methodology, Formal analysis. Sen Zhong: Software, Investigation, Data curation. Bowen Zheng: Investigation, Data curation. Diya Liu: Investigation, Data curation. Yuying Wang: Investigation, Funding acquisition, Data curation. Bolin Yu: Investigation, Funding acquisition, Data curation. Lin Fang: Supervision, Resources, Project administration, Funding acquisition, Conceptualization, Investigation, Funding acquisition, Data curation. Fengyuan Qian and Sen Zhong authors contributed equally to this work. All authors gave final approval and agree to be accountable for all aspects of the work.
Declaration of Competing Interest
The authors declare no conflicts of interest.
- Li YZ, Zheng JX, Deng YJ, Deng XY, Lou WY, Wei BJ, et al. (2022) Global Burden of Female Breast Cancer: Age-Period-Cohort Analysis of Incidence Trends From 1990 to 2019 and Forecasts for 2035. Frontiers in Oncology. 12.
- Dizon DS, Kamal AH (2024) Cancer statistics 2024: All hands on deck. Ca-a Cancer Journal for Clinicians. 74: 8-9.
- Cao W, Chen HD, Yu YW, Li N, Chen WQ (2021) Changing profiles of cancer burden worldwide and in China: a secondary analysis of the global cancer statistics 2020. Chinese Medical Journal. 134: 783-91.
- Koboldt DC, Fulton RS, McLellan MD, Schmidt H, Kalicki-Veizer J, McMichael JF, et al. (2012) Comprehensive molecular portraits of human breast tumours. Nature. 490: 61-70.
- Aftimos P, Oliveira M, Irrthum A, Fumagalli D, Sotiriou C, Gal-Yam EN, et al. (2021) Genomic and Transcriptomic Analyses of Breast Cancer Primaries and Matched Metastases in AURORA, the Breast International Group (BIG) Molecular Screening Initiative. Cancer Discovery. 11: 2796-811.
- Zhang YS, Chan HL, Garcia-Martinez L, Karl DL, Weich N, Slingerland JM, et al. (2020) Estrogen induces dynamic ERα and RING1B recruitment to control gene and enhancer activities in luminal breast cancer. Science Advances. 6.
- Leopold AV, Chernov KG, Verkhusha VV (2018) Optogenetically controlled protein kinases for regulation of cellular signaling. Chemical Society Reviews. 47: 2454-84.
- Gutierrez-Castellanos N, Da Silva-Matos CM, Zhou KK, Canto CB, Renner MC, Koene LMC, et al. (2017) Motor Learning Requires Purkinje Cell Synaptic Potentiation through Activation of AMPA-Receptor Subunit GluA3. Neuron. 93: 409-24.
- Norel X, Sugimoto Y, Ozen G, Abdelazeem H, Amgoud Y, Bouhadoun A, et al. (2020) International Union of Basic and Clinical Pharmacology. CIX. Differences and Similarities between Human and Rodent Prostaglandin E2 Receptors (EP1-4) and Prostacyclin Receptor (IP): Specific Roles in Pathophysiologic Conditions. Pharmacological Reviews. 72: 910-68.
- Yeh E, Dao DQ, Wu ZY, Kandalam SM, Camacho FM, Tom C, et al. (2018) Patient-derived iPSCs show premature neural differentiation and neuron type-specific phenotypes relevant to neurodevelopment. Molecular Psychiatry. 23: 1687-98.
- Farrow B, Rychahou P, Murillo C, O'Connor KL, Iwamura T, Evers BM (2003) Inhibition of pancreatic cancer cell growth and induction of apoptosis with novel therapies directed against protein kinase A. Surgery. 134: 197-205.
- Thomas J, Vanpatten SM, Howard P, Day KH, Mitchell RD, Sosnick T, et al. (1991) Expression In Escherichia-Coli and Characterization of the Heat-Stable Inhibitor of the Camp-Dependent Protein-Kinase. Journal of Biological Chemistry. 266: 10906-11.
- Collins SP, Uhler MD (1997) Characterization of PKI gamma, a novel isoform of the protein kinase inhibitor of cAMP-dependent protein kinase. Journal of Biological Chemistry. 272: 18169-78.
- Gamm DM, Uhler MD. ISOFORM-Specific Differences In The Potencies of Murine Protein-Kinase Inhibitors are Due to Nonconserved Amino-Terminal Residues. Journal of Biological Chemistry. 270: 7227-32.
- Scott JD, Fischer EH, Takio K, Demaille JG, Krebs EG (1985) Amino-Acid Sequence of the Heat-Stable Inhibitor of the Camp-Dependent Protein-Kinase from Rabbit Skeletal-Muscle. Proceedings of the National Academy of Sciences of the United States of America. 82: 5732-6.
- Dabanaka K, Chung S, Nakagawa H, Nakamura Y, Okabayashi T, Sugimoto T, et al. (2012) PKIB expression strongly correlated with phosphorylated Akt expression in breast cancers and also with triple-negative breast cancer subtype. Medical Molecular Morphology. 45: 229-33.
- VanPatten SM, Donaldson LF, McGuinness MP, Kumar P, Alizadeh A, Griswold MD, et al. (1997) Specific testicular cellular localization and hormonal regulation of the PKI alpha and PKI beta isoforms of the inhibitor protein of the cAMP-dependent protein kinase. Journal of Biological Chemistry. 272: 20021-9.
- Hauer JA, Barthe P, Taylor SS, Parello J, Padilla A (1999) Two well-defined motifs in the cAMP-dependermt protein kinase inhibitor (PKIα) correlate with inhibitory and nuclear export function. Protein Science. 8: 545-53.
- Kovacs Z, Schacht T, Herrmann AK, Albrecht P, Lefkimmiatis K, Methner A (2014) Protein kinase inhibitor β enhances the constitutive activity of G-protein-coupled zinc receptor GPR39. Biochemical Journal. 462: 125-32.
- Lum H, Hao ZP, Gayle D, Kumar P, Patterson CE, Uhler MD (2002) Vascular endothelial cells express isoforms of protein kinase A inhibitor. American Journal of Physiology-Cell Physiology. 282: C59-66.
- Chung S, Furihata M, Tamura K, Uemura M, Daigo Y, Nasu Y, et al. (2009) Overexpressing PKIB in prostate cancer promotes its aggressiveness by linking between PKA and Akt pathways. Oncogene. 28: 2849-59.
- Wan RX, Yang G, Liu QZ, Fu XK, Liu ZP, Miao HL, et al. (2022) PKIB involved in the metastasis and survival of osteosarcoma. Frontiers in Oncology. 12.
- Lacalamita A, Piccinno E, Scalavino V, Bellotti R, Giannelli G, Serino G (2021) A Gene-Based Machine Learning Classifier Associated to the Colorectal Adenoma-Carcinoma Sequence. Biomedicines. 9.
- Dou P, Zhang D, Cheng Z, Zhou G, Zhang L (2016) PKIB promotes cell proliferation and the invasion-metastasis cascade through the PI3K/Akt pathway in NSCLC cells. Exp Biol Med (Maywood). 241: 1911-8.
- Zhang JB, Song W, Wang YY, Liu MG, Sun MM, Liu H (2017) Study on correlation between PKIB and pAkt expression in breast cancer tissues. Eur Rev Med Pharmacol Sci. 21: 1264-9.
- Liu JF, Lichtenberg T, Hoadley KA, Poisson LM, Lazar AJ, Cherniack AD, et al. (2018) An Integrated TCGA Pan-Cancer Clinical Data Resource to Drive High-Quality Survival Outcome Analytics. Cell. 173: 400.
- Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology. 15.
- Hänzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-Seq data. Bmc Bioinformatics. 14.
- Bindea G, Mlecnik B, Tosolini M, Kirilovsky A, Waldner M, Obenauf AC, et al. (2013) Spatiotemporal Dynamics of Intratumoral Immune Cells Reveal the Immune Landscape in Human Cancer. Immunity. 39: 782-95.
- Haines CN, Klingensmith HD, Komara M, Burd CJ (2020) GREB1 regulates PI3K/Akt signaling to control hormone-sensitive breast cancer proliferation. Carcinogenesis. 41: 1660-70.
- Zhang P, Li X, Pan CW, Zheng XM, Hu BH, Xie RH, et al. (2022) Single-cell RNA sequencing to track novel perspectives in HSC heterogeneity. Stem Cell Research & Therapy. 13.
- Yuan HT, Yan M, Zhang GX, Liu W, Deng CY, Liao GM, et al. (2019) CancerSEA: a cancer single-cell state atlas. Nucleic Acids Research. 47: D900-D8.
- Prenzel T, Kramer F, Bedi U, Nagarajan S, Beissbarth T, Johnsen SA (2012) Cohesin is required for expression of the estrogen receptor-alpha gene. Epigenetics & Chromatin. 5.
- Senn HJ, Thürlimann B, Costa A, Cardoso F (2023) Future potential targets of antibody-drug conjugates in breast cancer (vol 69, pg 312, 2023). Breast. 69: 1.
- Musket A, Moorman JP, Zhang J, Jiang Y (2024) PKIB, a Novel Target for Cancer Therapy. Int J Mol Sci. 25.
- Dieci MV, Miglietta F, Guarneri V (2021) Immune Infiltrates in Breast Cancer: Recent Updates and Clinical Implications. Cells. 10.
- Wang D, Wu X, Sun Y (2022) Therapeutic targets and biomarkers of tumor immunotherapy: response versus non-response. Signal Transduction and Targeted Therapy. 7: 3784-810.
- Xiang X, Wang J, Lu D, Xu X (2021) Targeting tumor-associated macrophages to synergize tumor immunotherapy. Signal Transduction and Targeted Therapy. 6: 795-806.
- Togashi Y, Shitara K, Nishikawa H (2019) Regulatory T cells in cancer immunosuppression - implications for anticancer therapy. Nature Reviews Clinical Oncology. 16: 356-71.
- Sun YL, Yang J, Chen YC, Guo YD, Xiong J, Guo XQ, et al. (2024) PD-L2 Expression in Breast Cancer Promotes Tumor Development and Progression. Journal of Immunology Research. 2024.
- Yi M, Zheng XL, Niu MK, Zhu SL, Ge H, Wu KM (2022) Combination strategies with PD-1/PD-L1 blockade: current advances and future directions. Molecular Cancer. 21.

Figures at a glance