Exploration of the Potential Mechanism of Galangin for the Treatment of Pancreatic Cancer Based on Network Pharmacology and In Vitro Validation
AFFILIATIONS
AFFILIATIONS
AFFILIATIONS
AFFILIATIONS
AFFILIATIONS
Corresponding author (Address):
Received Date: March 08, 2025 Accepted Date: April 08, 2025 Published Date: April 11, 2025
doi:10.17303/jcrto.2025.13.104
Citation: Chen Mengchi, Zhang Jingfeng, Zheng Changlong, Liu Jianghua, Zhang Yongbiao (2025) Exploration of the Potential Mechanism of Galangin for the Treatment of Pancreatic Cancer Based on Network Pharmacology and In Vitro Validation. J Cancer Res Therap Oncol 13: 1-15
Abstract
Background: Galangin, a natural flavonoid, has been shown to have powerful anti-tumor effects of promoting apoptosis and autophagy, and inhibiting proliferation and metastasis. However, the role and mechanism of this natural compound on PC cell proliferation and migration is unknown. Therefore, this study aimed to investigate the potential mechanism of galangin for the treatment of pancreatic cancer in vitro on the basic of network pharmacology.
Methods: Swiss Target Prediction, Gene Cards, Drug Bank, STRING and other databases and software such as R language and Cytoscape 3.6.1 were used for target screening, gene annotation, GO and KEGG enrichment analysis, and construction of protein interaction network. The effects of galangin on the proliferation and migration of BxPC-3 pancreatic cancer cells were investigated in vitro using the CCK8 method, plate cloning, scratching, and Transwell method.; Finally, this study also validated important genes and pathways by molecular docking, GEPIA, HPA, and Western Blot. Results: A total of 98 signaling pathways were enriched by KEGG involving multiple pathways. Molecular docking results showed that galangin binds freely to the top 5 degree-ranked core targets. Galangin inhibited the proliferation and migration behavior of BxPC-3 pancreatic cancer cells in vitro. GEPIA, HPA, and Western Blot results further indicated that SRC and PTPN11 are important genes among the five core targets of galangin, and PI3K-Akt is an important signaling pathway in the enrichment pathway.
Conclusions: Galangin inhibits the proliferation and migration of BxPC-3 pancreatic cancer cells effectively, and its mechanism is closely related to the PI3K-AKT signaling pathway.
Keywords: Galangin; Pancreatic Cancer; Network Pharmacology; Mechanism of Action
Introduction
Pancreatic cancer(PC) is a malignancy extremely difficult to treat [1], has an increasing incidence and a high mortality rate owing to its ability to metastasize fast and the delay in diagnosis that often occurs [2-4]. A statistic from 2021 showed that in the United States, patients with PC have ranked the 4th highest mortality rate among all patients with cancer [5,6]. PC treatment include surgery, radiation, chemotherapy and so on. But these strategies are poor efficacy, and currently only adjuvant treatments are used to improve the quality of health and increase the survival rate of patients. Chinese herbal monomers are derived from medicinal plants and have the characteristics of multi-target and multi-channel regulation. Galangin as one of the Chinese herbal monomers, is a flavonoid extracted from the plant Alpinia officinarum Hance, which has various effects such as antioxidant and anti-inflammatory activity [7]. Additionally, galangin has demonstrated anti-tumor effects against cancers such as gastric and liver cancer [8,9], etc. However, there is little research on the therapeutic effects of this natural compound on PC. The proliferation and migration characteristics of tumor cells often cause tumor deterioration, which is an important reason for the rapid progression of tumor diseases. This study will focus on the anti-proliferation and anti-migration effects of galangin in pancreatic cancer.
Network pharmacology is a new technology that integrates pharmacology, bioinformatics and computer technology [10]. Using network pharmacology, we can effectively screen relevant target proteins and explain the various mechanism of action of drugs, this method being important for the study of plant-derived active ingredients.
In this study, the effects of galangin on the proliferation and migratory behavior of BxPC-3 PC cells were investigated in in vitro experiments. Using a network pharmacology approach, we further investigated the potential targets and mechanisms underlying the anticancer action of galangin in BxPC-3 PC cells. Thus, we aimed to provide a scientific basis for the use of galangin in the treatment of PC.
Methods
Cell Line and Culture
Human BxPC-3 PC cell (Procell, Wuhan, Hubei, China) line was inoculated in cell culture flasks and cultured with RPMI1640 (Gibco, USA) medium containing 10% fetal bovine serum (Procell, Wuhan, Hubei, China), 1% penicillin and streptomycin (Service bio, Wuhan, Hubei, China), and incubated continuously at 37 ℃, 5% CO2 and saturated humidity.
Determination of the Effect of Galangin on Cancer Cell Proliferation
The Cell Counting Kit 8 (CCK-8) (Lanjieke, Beijing, China) method was used to detect the effect of different concentrations of galangin (purity >98%, lot no.: MUST-21061401, Mansit, Chengdu, Sichuan, China) on the proliferation ability of BxPC-3 PC cells at different time points. First, BxPC-3 PC cells at logarithmic growth stage were seeded in 96-well culture plates at a density of 4×103 cells/well (100 μL/well); DMSO was added to obtain a high-concentration master batch which was stored at −20 ℃. according to the requirements of the instruction of galangin. Then diluted into 5, 10, 20, 40, 80 and 120 μmol/L of galangin solution respectively according to the experimental design. The cells were incubated for 24 h followed by the addition of 0.1% DMSO and each of the above concentration gradients of galangin-containing medium for interventions of 24, 48 and 72 h. Then 10 μL of CCK-8 solution were added to each well and cell viability was detected by measuring the optical density values at 450 nm in relation to an enzymatic standard after 1-4 h of incubation. For comparison, we used a blank control group and a 0.1% DMSO group. Five replicates were prepared for each group.
Plate Cloning Assay
BxPC-3 cells in the logarithmic growth phase were inoculated in 6-well plates (1 ml/well) at a density of 1×103 cells/well. Complete medium (control group), 20, 40 and 80 μmol/L galangin-containing medium were added 24 h after cell apposition growth. The duration of the exposure was 14 days, during which the fresh medium was replaced every 2 days. The cells were fixed and stained after clonal blocks visible to the naked eye appeared in the six-well plates. The number of clones was counted and photographed, and the average value was used to calculate the clone formation rate, as follows: Clone formation rate (%) = (number of clones/number of inoculated cells) × 100.
Scratching Assay
BxPC-3 cells at logarithmic growth stage were inoculated in 6-well plates and incubated for 24 h in an incubator at 37 ℃, 5% CO2 and saturated humidity. The cells on the bottom of the culture plate were scratched perpendicularly with a 200 μL yellow pipette tip and washed twice with PBS. The control group was treated with serum-free medium, while the experimental group was treated with serum-free medium containing different concentrations of galangin (20, 40, 80 μmol/L). Photographs were taken at 0 h, 24 h and 48 h in the same positions. Scratch widths were measured using Image J software to calculate cell migration rates. The rate of mobility was calculated using the following formula: Mobility %= (W0h-Wnh)/W0h, W0h is scratch width at 0 h, Wnh is scratch width after n h.
Transwell Assay
Matrix gel was added to the bottom membrane of the transwell upper chamber and incubated overnight at 37 °C. Bxpc-3 cells were cultured without serum for 24 h, detached from the culture plate with trypsin, re-suspended in serum-free culture medium, inoculated in Transwell upper chamber (at a density of 1×105 cells/well). Drug-containing medium - 15% fetal bovine serum containing galangin at various concentrations (20, 40, 80 μmol/L), was added in the lower chamber. The transwell chambers were incubated at 37 °C for 24 h and then removed. The stromal gel was removed from the upper chamber with a cotton swab, the cells were fixed with 5% paraformaldehyde for 20 min before being stained with 1% crystal violet for 5-10 min and photographed under a microscope. The results were analyzed using Image J software.
Candidate Targets of Galangin
We entered the search term "galangin" in PubChem database (https://pubchem.ncbi.nlm.nih.gov/), checked its 2D structure, obtained the corresponding Canonical SMILES, and imported them into SwissTargetPrediction (http://www.swisstargetprediction.ch/), SEA (https://sea.bkslab.org/) and Drug Bank (https://go.drugbank.com/) databases. After screening, the corresponding targets were obtained.
Candidate Target Genes for PC
Using "pancreatic cancer" as the search term, we acquired PC-related targets by retrieving TTD (http://db.idrblab.net/ttd/), OMIM (https://omim.org/), Gene Cards (https://www.genecards.org/), Drug Bank (https://go.drugbank.com/), and Pharm GKB (https://www.pharmgkb.org/) databases. and to obtain concatenation and delete duplicate target genes.
Screening of Core Targets and Construction of Protein Interactions Network
A common target is obtained by intersection of drug and disease target genes. The results were imported into STRING database [11] (https://www.string-db.org/), the species was limited to "Homo sapiens" and "minimum required interaction score" parameter was set to >0.9 (highest confidence). The rest of the parameters were set to the default settings on the website to predict the functional interactions of target-related proteins. Cytoscape software v. 3.6.1 was used to build a protein-protein interaction (PPI) network, and the top 5 hub genes were filtered using the degree algorithm in cytoHubba plugin.
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) Functional Enrichment Analysis
Using the "clusterProfiler" package of R and setting the statistical significance at P-value < 0.05, we analyzed the PC-related targets of galangin for biological process (BP), cellular component (CC) and molecular function (MF). The results were analyzed and visualized using the R packages "enrichplot", "ggplot2" and "circlize".
Construction of “Drug-Target-Signaling Pathway” Network
To understand the relationship between the targets of galangin, PC and the corresponding signaling pathways, a "drug-target-signaling pathway" network was constructed using Cytoscape 3.6.1 software.
Molecular Docking
The top 5 key genes were selected for molecular docking with galangin to gain a preliminary understanding of the therapeutic mechanism of the latter. The files containing the protein structure corresponding to the above key genes were downloaded from Protein Data Bank (PDB) database (https://www.rcsb.org/) in PDB format, and imported into Auto dock 1.5.7 software for molecular docking with galangin. A docking score of less than -5 kcal/mol indicated that target protein/galangin can form a complex [12], which was visualized using Pymol 2.5.2 software.
Differential expression of key genes
The top 5 key genes were imported into the online website GEPIA (http://gepia.cancer-pku.cn/) to verify their mRNA expression levels in human PC tissues in The Cancer Genome Atlas (TCGA) and Genotype-Tissue Expression (GTEx) databases [13]; additionally, they were also imported in the Human Protein Atlas (HPA) database (https://www. proteinatlas.org/) for retrieving the protein expression of key genes [14].
Western Blot
Bxpc-3 PC cells at logarithmic growth stage were inoculated in 6-well plates, and when the cell density was about 70%-80%, the control and experimental groups were cultured for 48 h with complete medium and 80 μmol/L of galangin-containing medium, respectively. After 48 h, the cells were washed twice with PBS, and 200 μL of lysis solution was added to each well. Cells were further lysed on ice for 30 min, the suspension was centrifuged at 4 °C and 12,000 rpm/min for 5 min and the supernatant was collected. After the protein concentration was determined using bicinchoninic acid (BCA) method, a protein solution was added to the loading buffer at a ratio 4:1 protein/loading buffer, mixed and denatured at 100 °C for 10 min using a PCR instrument. After denaturation, the protein was stored at -20 °C. After 60V, 25min to 120V, 65min constant voltage electrophoresis, 200mA constant current transfer membrane for 70min, and 5% nonfat milk powder for 1h blocking, the diluted primary antibody SRC (1:800, Sanying, Wuhan, Hubei, China), PIK3R1(1:2,0000, Sanying, Wuhan, Hubei, China), PTPN11(1:2,000, Sanying, Wuhan, Hubei, China), AKT(1:10000, Abcam, Cambridge, UK), E-cadherin(1:10000, Abcam, Cambridge, UK), GAPDH(1:20,000, Sanying, Wuhan, Hubei, China) was added and incubated overnight at 4 °C. The membrane was washed 3 times with Tris-buffered saline and Tween (TBST) for 5 min each time, the diluted secondary antibody was added, the mix was incubated at room temperature for 1 h, the membrane was again washed 3 times with TBST. The ECL mix was prepared according to the instructions of the manufacturer, added to the membrane which was further incubated for 5 min. The grey scale values of each band were analyzed using Image J software.
Statistical Analysis
SPSS 23.0 software was used for data analysis. Data was expressed as mean ± standard deviation (X±s). One-way ANOVA was used for comparison between multiple groups, Fisher’s least significance difference test (LSD-t) was used for two-way comparison between groups if the variance was the same, and Tambane's T2 test was used for two-way comparison between groups if the variance was not similar. The difference of P < 0.05 means statistical significance.
Results
Effect of Galangin on the Proliferation of BxPC-3 PC Cells
The experimental results of CCK-8 method for assessing the proliferation of BxPC-3 cells (Figure 1A) showed that galangin inhibits the proliferation of BxPC-3 cells at 24, 48 and 72 h, in a time- and concentration-dependent manner, i.e., the proliferation of cancer cells decreased with the increase of the galangin concentration and time of exposure. Compared with control, the most significant inhibition of cell proliferation was observed for the 80 μmol/L concentration of galangin at 24, 48, and 72 h(P<0.0001).
Plate Cloning Assay to Detect the Effect of Galangin on the Independent Proliferation of BxPC-3 PC Cells
The experimental results (Figure 1B) showed that the clone formation rate of BxPC-3 cells was significantly lower in the 20, 40 and 80 μmol/L galangin groups than in the control group, and the effect was dose-dependent.
Effect of Galangin on the Migratory Behavior of BxPC-3 PC Cells
The results of the scratch assay (Figure 2) showed that the migratory capacity of Bxpc-3 cells after 24 h of exposure to galangin 80 μmol/L was lower than that of the control. After 48 h of exposure to galangin, compared with control, the migratory capacity of Bxpc-3 cells was significantly reduced in the 20, 40, and 80 μmol/L groups. The results of the transwell assay (Figure 3) showed that 20, 40 and 80 μmol/L galangin-containing medium interfered with Bxpc-3 cells migration following 24 h of exposure. Compared with control, the number of cells penetrating the artificial basement membrane decreased following galangin treatment in a concentration-dependent manner. The results indicated that galangin inhibits the migration of PC cells.
Acquisition of Galangin’s Structure and its Targets of Action
The 2D structure of Galangin and the structural formula of Canonical SMILES were obtained from PubChem database (Figure 4). In total, 144 relevant targets were obtained from SwissTargetPrediction, SEA and DrugBank databases.
PC-Associated Target Genes
In total, 4146 PC-related target genes were screened in TTD, OMIM, GeneCards, DrugBank and PharmGKB databases.
Construction of Protein Interaction Network and Screening of Core Targets
Using the analysis parameters above-mentioned, we obtained a total of 82 nodes (Figure 5A). The average node degree was 3.1 and the PPI enrichment value was <1.0e-16. The results were imported into Cytoscape 3.6.1 to remove the unconnected target proteins and construct the main target protein interactions network between galangin and PC (Figure 5B). The top 5 hub genes were identified as SRC, PIK3R1, PTPN11, VEGFA, EGFR (Figure 5C).
Results of GO and KEGG Enrichment Analysis
The GO annotation analysis and KEGG signaling pathway enrichment analysis were performed on 82 PC-related targets of galangin, and a total of 1112 biological processes, 41 cellular components, 122 molecular functions and 98 KEGG signaling pathways were obtained. In the GO annotation analysis (Figure 6A), the most significant biological processes involved were protein serine/threonine kinase activity, protein complex kinase activity, tetrapyrrole binding, etc; the most significant cellular components included chromosomal regions, membrane microregions, membrane rafts, etc; the most significant molecular functions included response to oxidative stress, cellular response to chemical stress, cellular response to oxidative stress, etc. The KEGG signaling pathway analysis (Figure 6B) showed that the top 30 enriched pathways in PC are: PI3K-Akt, Ras, MAPK, Rap1, IL-17, AMPK, and VEGF signaling pathways, etc. Among them, PI3K-Akt signaling pathway was most significantly enriched.
"Drug-target-signaling pathway" Network
The potential targets and major signaling pathways modulated by galangin and related to PC were introduced into Cytoscape 3.6.1 software to construct a "drug-target-signaling pathway" network (Figure 7). The results showed that galangin could act against PC through multiple targets involved in different signaling pathways.
Molecular Docking Results
As shown in the figure (Figure 8), five key genes were able to bind galangin freely and the docking energy of each gene was less than -5 kcal/mol: SRC (PDB ID: 4M4Z): -5.58 kcal/mol; PIK3R1 (PDB ID: 2IUG): -5.33 kcal/mol; PTPN11 (PDB ID: 4RDD): -5.59 kcal/mol; VEGFA (PDB ID: 2VGH): -5.11 kcal/mol; EGFR (PDB ID: 4LLO): -8.7 kcal/mol. The molecular docking results provide preliminary validation that galangin can combine with key targets related to PC to exert pharmacological effects.
Differences in Expression of Key Genes Data collected from GEPIA database indicated that the levels of mRNA expression of all five target genes are higher in human PC tissues than in normal tissues (Figure 9A). However, only the increase in the expressions of SRC, PIK3R1, and PTPN11 genes reaches statistical significance. In the HPA database, the expression levels of key target proteins encoded by SRC, PIK3R1, PTPN11, VEGFA, EGFR were higher in human PC tissues than in normal tissues (Figure 9B).
Differences in Expression of Key Genes
Data collected from GEPIA database indicated that the levels of mRNA expression of all five target genes are higher in human PC tissues than in normal tissues (Figure 9A). However, only the increase in the expressions of SRC, PIK3R1, and PTPN11 genes reaches statistical significance. In the HPA database, the expression levels of key target proteins encoded by SRC, PIK3R1, PTPN11, VEGFA, EGFR were higher in human PC tissues than in normal tissues (Figure 9B).
Western Blot
The results of the protein blotting assay (Figure 10) showed that the proteins expressions of SRC, PI3KR1, PTPN11, E-cadherin, Akt were significantly reduced after exposure of BxPC-3 PC cells to 80 μmol/L galangin compared with the control (all P<0.001).
Discussion
Tumors exhibit various behaviors during growth and development, in terms of proliferation, induction of angiogenesis, activation of invasion and metastasis signals, etc. Proliferation, migration and invasion, are essential traits of tumors with high malignancy, and, as such, are important biological characteristics of PC [15]. Therefore, studying their patterns in this type of cancer and their related mechanisms can help us identify a more effective treatment of PC. In this study, it was confirmed by in vitro experiments of CCK8, plate cloning, scratch, and Transwell, etc that galangin had an inhibitory effect on the proliferation and migration of BxPC-3 PC cells.
Galangin belongs to the group of flavonoids extracted from medicinal plants. The efficacy of various flavonoids against PC, including kaempferol, apigenin and quercetin, etc, has been widely investigated. Jungwhoi Lee [16] showed that kaempferol inhibits PC cell migration by blocking EGFR-related Src, AKT and ERK1/2 signaling pathways, etc. Lei Cao [17] showed that resveratrol inhibited high glucose-derived ROS-induced invasion and migration of PC cells by inhibiting ERK and p38 MAPK signaling pathways. Dingguo Wu [18] confirmed that apigenin produced growth inhibition in PC cells by inhibiting IKK-β-mediated NF-κB pathway. Yangyang Guo [19] showed that quercetin inhibited the proliferation, migration and invasion of pancreatic ductal adenocarcinoma by inhibiting SHH and TGF-β/Smad signaling pathways. Thus, it can be concluded PC is regulated by multiple targets and signaling pathways. The mechanisms of action of flavonoids, including galangin, are complex, therefore understanding the targets and underlying mechanisms of galangin’s anticancer effect requires network pharmacology to be combined with other related methods.
In this study, employing a network pharmacology approach, 82 PC-related targets of galangin were identified using relevant databases. the most significant five core targets were obtained by cytoHubba's degree algorithm, namely SRC, PIK3R1, PTPN11, VEGFA and EGFR. Among them, SRC, PIK3R1 and PTPN11 encode proteins that are cell signaling regulators; VEGFA and EGFR encode tumor vascular growth factors; SRC encodes a non-receptor tyrosine kinase, which plays an important role in cell surface and transmembrane protein signaling in tumors, and also participates in the activation of several downstream signaling pathways [20,21]. PIK3R1 encodes the regulatory subunit p85α of PI3K, which is essential for the regulation of PI3K-Akt signaling pathway and inhibition of cancer cell proliferation and cell cycle progression [22]. PTPN11 is associated with the tyrosine phosphatase family and mainly encodes the tyrosine phosphatase Shp2, which is a key linkage point for various intracellular oncogenic signaling pathways, including PI3K/AKT, PD-1/PD-L1 and other pathways [23,24]. VEGFA is associated with tumor angiogenesis and is involved in regulating cancer cell proliferation, apoptosis, and vascular permeability through VEGF-A, which activates VEGFR2-mediated pathways including PI3K-Akt and ERK signaling pathways, etc [25,26]. EGFR is derived from ErbB family of receptor tyrosine kinases, which can regulate the tumor microenvironment through autophosphorylation and initiation of downstream signaling pathways, such as MAPK signaling pathway [27-29].
The results of molecular docking validated the hypothesis galangin binds to the protein expressions of the above-mentioned five core genes. In addition, we learned from the HPA database that the expression levels of the five key target proteins were increased in human PC tissues compared with normal pancreatic tissues, while data in the GEPIA database showed that only SRC, PIK3R1, and PTPN11 had significantly higher levels of expression in human PC tissues than in normal ones; considering the different data sources of HPA and GEPIA databases and the differences in the results provided, as well as the limitations of molecular docking, to further validate our hypothesis, we selected SRC, PIK3R1 and PTPN11 for western Blot. The results showed that compared with the control group, the expression of SRC, PTPN11 and PIK3R1 related-proteins decreased significantly after exposure to galangin at a concentration of 80 μmol/L. Here, we further confirm SRC, PTPN11 and PIK3R1 as important targets of galangin. KEGG enrichment results showed that multiple pathways were involved, including the PI3K-Akt, MAPK, and VEGF signaling pathways.etc. Interestingly, we performed in vitro experiments to validate the PI3K-Akt signalling pathway, which is the most significantly enriched by KEGG, and showed that PI3K p85 and Akt proteins were inhibited after galangin intervention compared to the control group. We speculate that galangin may regulate pancreatic cancer cell proliferation and invasion by acting on the PI3K-Akt signaling pathway.
In summary, galangin can effectively inhibit the proliferation and migration of BxPC-3 pancreatic cancer cells, and its mechanism is closely related to PI3K-AKT signaling pathway.
Acknowledgments
None
Funding
None
Availability of Data and Materials
All data generated and analyzed during the present study are included in this article.
Ethics Approval and Consent to Participate
Not applicable.
Patient Consent for Publication
Not applicable.
Authors’ Contribution
Chen Mengchi and Zhang Jingfeng have made equal contributions to this paper: project design, experimental design and implementation, writing – original draft, writing - review and editing. Zheng Changlong: Chart design and revision. Liu Jianghua and Zhang Yongbiao: Conception, communication and revision.
Disclosure Statement
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
- NE Cortez, GG Mackenzie (2021) Ketogenic Diets in Pancreatic Cancer and Associated Cachexia: Cellular Mechanisms and Clinical Perspectives, Nutrients, 13: 3202.
- P Rawla, T Sunkara, V Gaduputi (2019) Epidemiology of Pancreatic Cancer: Global Trends, Etiology and Risk Factors, World J Oncol. 10: 10-27.
- H Sung, J Ferlay, RL Siegel, M Laversanne, I Soerjomataram, A Jemal, F Bray (2021) Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries, CA Cancer J Clin. 71: 209-49.
- R Ranganath, Q Chu (2021) Global trends in pancreas cancer among Asia-Pacific population, J Gastrointest Oncol. 12: S374-386.
- C.P.S.A. CSoSCMA (2021) Guidelines for the diagnosis and treatment of pancreatic cancer in China (2021), Zhonghua Wai Ke Za Zhi. 59: 561-77.
- J Cai, H Chen, M Lu, Y Zhang, B Lu, L You, T Zhang, M Dai, Y Zhao (2021) Advances in the epidemiology of pancreatic cancer: Trends, risk factors, screening, and prognosis, Cancer Lett. 520: 1-11.
- Y Xiong, X Lai, W Xiang, J Zhou, J Han, H Li, H Deng, L Liu, J Peng, L Chen (2020) Galangin (GLN) Suppresses Proliferation, Migration, and Invasion of Human Glioblastoma Cells by Targeting Skp2-Induced Epithelial-Mesenchymal Transition (EMT), Onco Targets Ther. 13: 9235-44.
- X Liang, P Wang, C Yang, F Huang, H Wu, H Shi, X Wu (2021) Galangin Inhibits Gastric Cancer Growth Through Enhancing STAT3 Mediated ROS Production, Front Pharmacol. 12: 646628.
- X Zhong, S Huang, D Liu, Z Jiang, Q Jin, C Li, L Da, Q Yao, D Wang (2020) Galangin promotes cell apoptosis through suppression of H19 expression in hepatocellular carcinoma cells, Cancer Med. 9: 5546-57.
- AL Hopkins, Network pharmacology, Nat Biotechnol. 25: 1110-1.
- D Szklarczyk, JH Morris, H Cook, M Kuhn, S Wyder, M Simonovic, et al. (2017) quality-controlled protein-protein association networks, made broadly accessible, Nucleic Acids Res. 45: D362-8.
- S Song, F Wen, S Gu, P Gu, W Huang, S Ruan, X Chen, et al. (2022) Network Pharmacology Study and Experimental Validation of Yiqi Huayu Decoction Inducing Ferroptosis in Gastric Cancer, Front Oncol. 12: 820059.
- Z Tang, C Li, B Kang, G Gao, C Li, Z Zhang (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses, Nucleic Acids Res. 45: W98-102.
- A Digre, C Lindskog (2021) The Human Protein Atlas-Spatial localization of the human proteome in health and disease, Protein Sci. 30: 218-33.
- D Hanahan, RA Weinberg (2011) Hallmarks of cancer: the next generation, Cell, 144: 646-74.
- J Lee, JH Kim (2016) Kaempferol Inhibits Pancreatic Cancer Cell Growth and Migration through the Blockade of EGFR-Related Pathway In Vitro, PLoS One, 11: e0155264.
- L Cao, X Chen, X Xiao, Q Ma, W Li (2016) Resveratrol inhibits hyperglycemia-driven ROS-induced invasion and migration of pancreatic cancer cells via suppression of the ERK and p38 MAPK signaling pathways, Int J Oncol. 49: 735-43.
- DG Wu, P Yu, JW Li, P Jiang, J Sun, HZ Wang, LD Zhang, MB Wen, P Bie (2014) Apigenin potentiates the growth inhibitory effects by IKK-β-mediated NF-κB activation in pancreatic cancer cells, Toxicol Lett. 224: 157-64.
- Y Guo, Y Tong, H Zhu, Y Xiao, H Guo, L Shang, W Zheng, S Ma, X Liu, Y Bai (2021) Quercetin suppresses pancreatic ductal adenocarcinoma progression via inhibition of SHH and TGF-β/Smad signaling pathways, Cell Biol Toxicol. 37: 479-96.
- V Vlaeminck-Guillem, G Gillet, R Rimokh (2014) SRC: marker or actor in prostate cancer aggressiveness, Front Oncol. 4: 222.
- Y Liu, M Dai, P Yang, L Cao, L Lu (2022) Src-homology domain 2 containing protein tyrosine phosphatase-1 (SHP-1) directly binds to proto-oncogene tyrosine-protein kinase Src (c-Src) and promotes the transcriptional activation of connexin 43 (Cx43), Bioengineered, 13: 13534-43.
- LW Cheung, GB Mills (2016) Targeting therapeutic liabilities engendered by PIK3R1 mutations for cancer treatment, Pharmacogenomics, 17: 297-307.
- X Zhao, S Hu, L Wang, Q Zhang, X Zhu, H Zhao, et al. (2014) Functional short tandem repeat polymorphism of PTPN11 and susceptibility to hepatocellular carcinoma in Chinese populations, PLoS One, 9: e106841.
- X Yuan, H Bu, J Zhou, CY Yang, H Zhang (2020) Recent Advances of SHP2 Inhibitors in Cancer Therapy: Current Development and Clinical Application, J Med Chem. 63: 11368-96.
- M. Shibuya (2013) Vascular endothelial growth factor and its receptor system: physiological functions in angiogenesis and pathological roles in various diseases, J Biochem. 153: 13-9.
- Y Zhang, RA Brekken (2022) Direct and indirect regulation of the tumor immune microenvironment by VEGF, J Leukoc Biol, 111: 1269-86.
- S Sigismund, D Avanzato, L Lanzetti (2018) Emerging functions of the EGFR in cancer, Mol Oncol. 12: 3-20.
- P Bhatia, V Sharma, O Alam, A Manaithiya, P Alam, Kahksha MT Alam, M Imran (2020) Novel quinazoline-based EGFR kinase inhibitors: A review focussing on SAR and molecular docking studies (2015-2019), Eur J Med Chem. 204: 112640.
- J Zhou, Q Ji, Q Li (2021) Resistance to anti-EGFR therapies in metastatic colorectal cancer: underlying mechanisms and reversal strategies, J Exp Clin Cancer Res. 40: 328.
FIGURE 1
Figure 1: Effect of galangin on the proliferation of BxPC-3 PC cells. (A) CCK-8; (B) Plate cloning assay. *P<0.05,***P<0.001, ****P<0.0001.
FIGURE 2
Figure 2: Scratching assay. nsP>0.05,*P<0.05,**P<0.01,***P<0.001.
FIGURE 3
Figure 3: Transwell assay. ***P<0.001,****P<0.0001.
FIGURE 4
Figure 4: 2D structure of Galangin and Canonical SMILES structural formula.
FIGURE 5
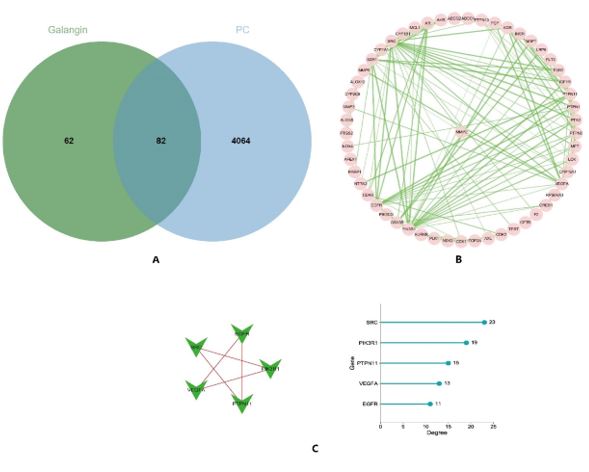
Figure 5: (A) The intersection is the target of Galangin action in pancreatic cancer; (B) Protein-protein interaction network (PPI); (C) Degree algorithm top 5 targets.
FIGURE 6
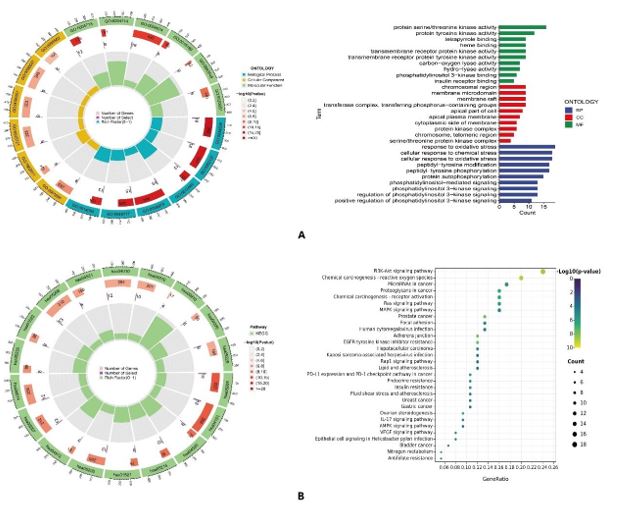
Figure 6: (A) GO annotation analysis; (B) KEGG pathway enrichment analysis.
FIGURE 7
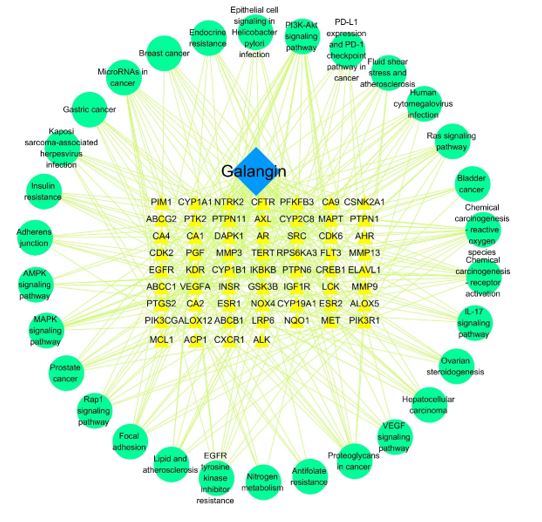
Figure 7: “Drug-target-signaling Pathway” Network.
FIGURE 8
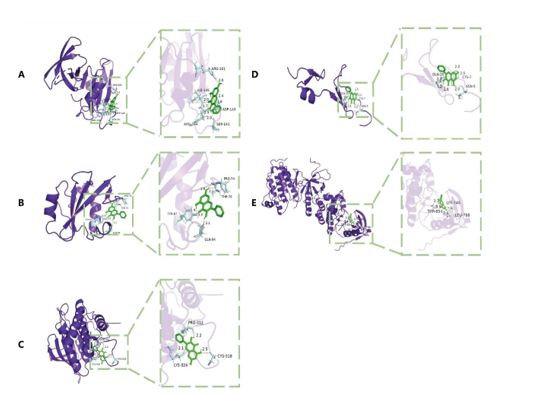
Figure 8: Schematic diagram of molecular docking. (A–E) represent the molecular binding of Galangin with SRC, PIK3R1, PTPN11, VEGFA, and EGFR, respectively.
FIGURE 9
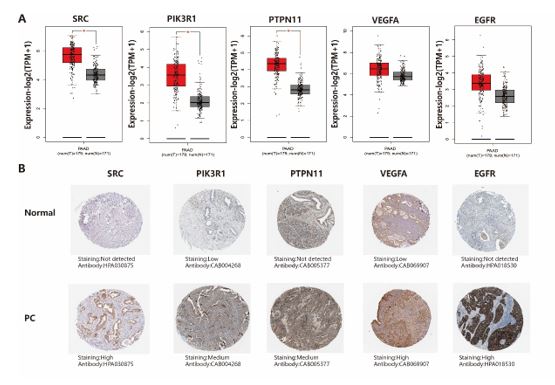
Figure 9: mRNA expression level in the GEPIA database and the protein expression levels in the HPA database. (A) Box plots showing the mRNA expression levels of SRC, PIK3R1, PTPN11, VEGFA, and EGFR. Red represents Tumor, Gray represents normal. (B) Protein expression levels of SRC, PIK3R1, PTPN11, VEGFA and EGFR in normal pancreatic tissue and pancreatic cancer tissue.
FIGURE 10
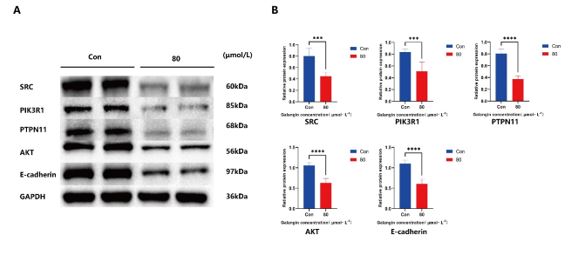
Figure 10: Effect of 80μmol/L galangin on SRC, PIK3R1, PTPN11, AKT, and E-cadherin protein. (A) Protein bands of SRC, PIK3R1, PTPN11, AKT and E-cadherin in control and intervention groups; (B) Relative protein expression of SRC, PIK3R1, PTPN11, AKT and E-cadherin. ***P<0.001,****P<0.0001.

Figures at a glance