Nutritional Value Analysis of the Larvae of the superworm Zophobas morio by using Proteomics Technology
Received Date: March 22, 2025 Accepted Date: April 22, 2025 Published Date: April 25, 2025
doi:10.17303/jfn.2025.11.201
Citation: Yu-Ting LIN, Hui-Juan YANG, Xin-Yan Li, Xiao-Ru JIAN, Yu-Huan HUANG, et al. (2025) Nutritional Value Analysis of the Larvae of the superworm Zophobas morio by using Proteomics Technology. J Food Nutr 11: 1-11
Abstract
The superworm Zophobas morio is becoming increasingly important food insects for people. In this paper, larvae of the superworm were used as research materials to detect and analyze the protein data of larvae of Z. morio by using data independent acquisition technology. Gene Ontology database, Kyoto Encyclopedia of Genes and Genomes database, clusters of euKaryotic Orthologous Groups database and other databases were used for protein identification and detection, i.e., proteomic analysis. The results showed that the number of detected proteins was 4589, and 91.9% of them were successfully annotated, among which Trx-2 protein was found to be a high-quality protein that can be absorbed by the human body and the larvae of Z. morio were developed in the food industry to grind insect powder as a food raw material, and the edible benefits of amino acid oral liquid manufacturing need to be further studied, so PCR amplification analysis was carried out for its Trx-2 gene.
Keywords: Zophobas morio; Larva; Proteomics; Immunoprotein; Trx-2 protein
Introduction
The superworm Zophobas morio is relatively high in proteins, amino acids, fatty acids, and trace elements [1]. In recent years, research on it has gradually shifted from traditional feed to edible nutrition research, demonstrating its potential as food and feed. There is comparatively less research on Z. morio at home and more research abroad, especially in the application of insects as food and feed. Although insects are still not widely used at home and abroad as a food source, studies have shown that the higher the consumer's awareness of insect nutrition, the higher their acceptance of raw insect food and the less resistant they are to fisheries and insect-fed livestock meat products. The market is not yet saturated and still needs to be expanded, resulting in a huge market for people's consumer demand, which needs to be further explored [2]. Regulations allow specific insect species to be used in aquafeeds and human food, and Z. morio larvae are also approved for use in human diets, providing a legal basis for the promotion of insect foods [3,4]. In summary, Z. morio has a promising application in the food and feed sector. With the deepening of research and the support of regulations, they may become an important source of nutrition in the future. Currently, there has been some progress in the study of the Z. morio transcriptome. Several studies have selected 11 transporters from the midgut transcriptome of Z. morio, revealing gene expression in different parts of the midgut and the physiological processes in which these transporters may be involved [5]. Some studies have also proposed antennal transcriptome sequencing information for Z. morio, which provides a solid foundation for future research on its chemo sensing genes [6].
In addition, proteomics studies can directly reveal the expression level, modification status, and function of proteins in organisms. Insect proteomics mainly focuses on the study of pathogenic mechanisms and differential analysis of specialized survival [9]. Zhao et al. identified differences in DEG expression patterns during the overwintering period of four western bees through transcriptomic identification and KEGG and GO enrichment analysis. The study explains the cold resistance mechanism of Western honey bees [7]. Guo et al. performed deep sequencing of the intestinal tract of Chinese honey bee larvae under coccal stress using proteomics and performed in-depth analysis using related software [8]. However, there is comparatively little research on the Z. morio proteome. Therefore, it is very necessary to conduct proteomic studies on the proteins of Z. morio, and the study of annotating protein functions in its proteomics is increasingly valuable. The preliminary statistical analysis of functional proteins in the proteome of Z. morio larvae was achieved by applying data independent acquisition (DIA) technology to analyze the proteome of Z. morio larvae to obtain the classification of annotated protein functions. PCR amplification analysis of the nutritionally representative immune protein Trx-2 in protein annotation. The results showed preliminary results that the larvae of Z. morio contained the Trx-2 protein. This finding has certain positive benefits for the subsequent breeding, food research and development, and market expansion of Z. morio larvae, and also lays a foundation for the nutritional analysis of subsequent Z. morio larvae.
Materials and Methods
Experimental Materials
The animal materials used in this experiment were 50 larvaes of Z. morio purchased online from Yangjiang, Guangdong in October 2023. These Z. morio larvaes were reared in the laboratory, and individuals exhibiting robust vitality were selected and promptly treated with liquid nitrogen for subsequent use.
Experimental Methods
Sample preprocessing includes protein extraction, denaturation, reduction and alkylation, enzymatic digestion, and peptide desalting. In this project, the iST sample preprocessing kit (PreOmics, Germany) was used for the preprocessing of tissue samples. The tissue samples were ground in liquid nitrogen, and an appropriate amount of sample was taken and mixed with 50 µl of lysis buffer. The mixture was heated at 95°C with 1000 rpm shaking for 10 minutes. After cooling to room temperature, trypsin digestion buffer was added, and the samples were incubated at 37°C with 500 rpm shaking for 2 hours. The enzymatic reaction was stopped by adding a termination buffer. The peptides were desalted using the iST cartridge provided in the kit, and eluted with 2 × 100 µl of elution buffer. The eluted peptides were vacuum-dried and stored at -80°C.
Construction Process of the Z. morio Larvae Protein
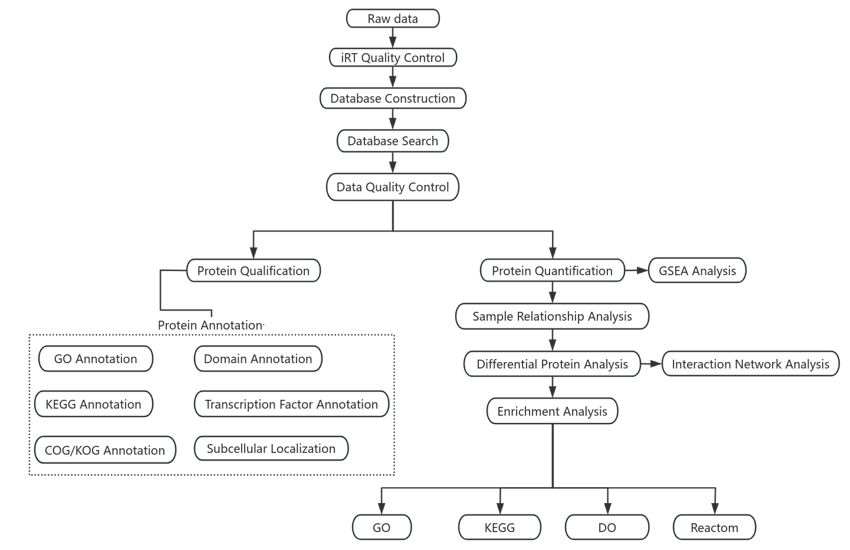
The proteome construction process is shown in Figure 1.
In this experiment, DIA technology was used for proteome data identification and annotation, and through the comparison of detection information with traditional databases, it had the advantages of high quantitative and qualitative accuracy, high repeatability, and data traceability. It is suitable for protein detection of large sample sizes and complex systems. Therefore, DIA technology can obtain more accurate and abundant proteome analysis results of the Z. morio larvae.
Results
Identification Results of Larval Protein of the Z. morio Larvae
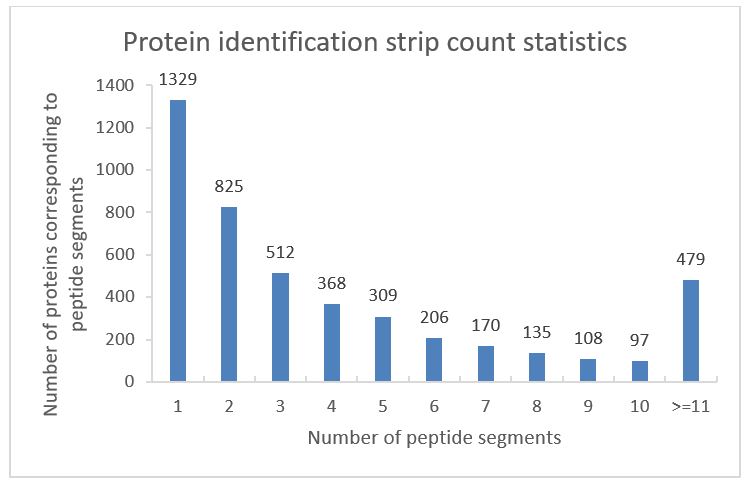
Proteins were extracted from the Z. morio larvae and enzymatically hydrolyzed. Before mass spectrometry detection, add Biognosys's internal standard reagent iRT Kit to each sample for the calibration of peptide retention times. Calibration is performed based on the retention time (RT) of peptides in chromatography, and use QuiC (Biognosys) software to perform quality control on raw mass spectrometry data, and assess whether the quality control indicators are similar across various samples. If the quality mark results are similar, it indicates that the test is well reproducible. Then, database construction of the collected MS data using Pulsar software, and the identification standards of Precursor Threshold 1.0% FDR and Protein Threshold 1.0% FDR were met at the peptide and protein levels, respectively. A total of 25571 protein peptides, including 23235 peptides segments, 4538 protein groups and 4589 proteins, were detected. Statistics on the results obtained (Table 1).
Annotation Results of the Z. morio Larvae Protein
To clearly measure the Z. morio larvae, protein domain alignment was performed by comparing with the GO, KEGG, and KOG databases. The total number of proteins was 4589, of which 4221 were successfully annotated in the databases. Specifically, 4058 proteins were annotated in the GO database, 2271 in the KEGG database, and 3505 in the KOG database. The effective protein function annotation rate was 91.98%.
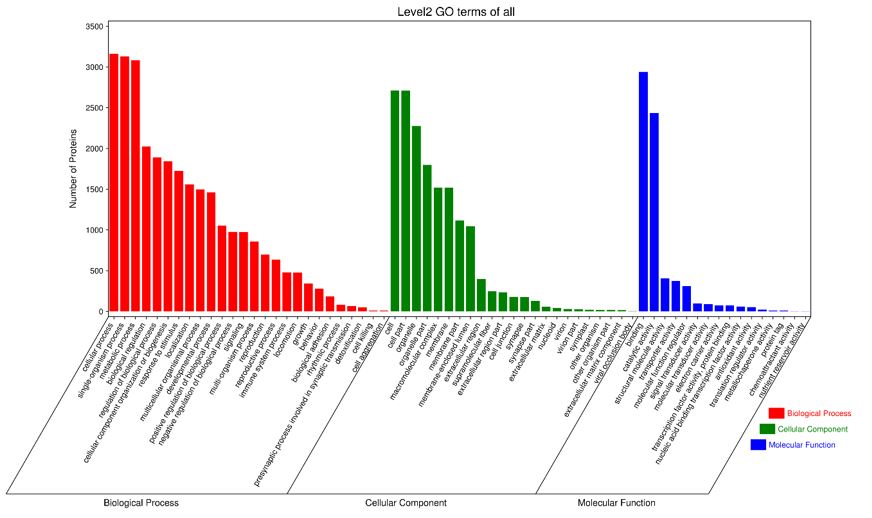
A total of 4058 proteins were annotated by GO function, and according to the GO enrichment table, most of the proteins were enriched in biological processes (Figure2). Among them, the functions of proteins in biological processes were divided into 26 detailed functions, of which 3160 were the most enriched proteins in cellular processes. According to the cellular component, the protein function can be divided into 21 categories, and the most enriched protein is the cell composition. According to the molecular function, it can be divided into 12 functions, and the function of enriching the most proteins is the binding function.
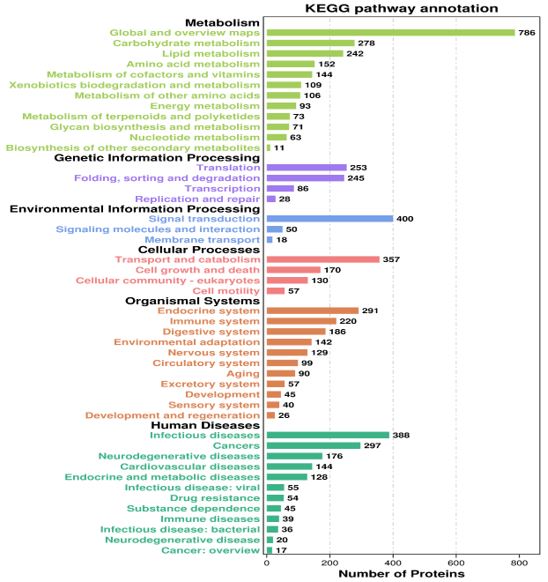
The most enriched pathway in KEGG is metabolism, with a total of 2271 proteins annotated to metabolic pathways. According to the analysis of the figure (Figure 3), these proteins can be divided into six branches, namely metabolism, genetic information processing, environmental information processing system, cellular processes, human diseases, and biological systems. The top 10 significantly enriched pathways have been listed and statistically analyzed (Table 2), Among them, three pathways are related to human diseases, two pathways are related to metabolism, genetic information, and cellular processes, while pathways related to biological systems appear only once. Among them, the metabolic pathway has the highest number of annotated proteins, with 784, the lowest is the endocytosis pathway, with only 89 annotated proteins.
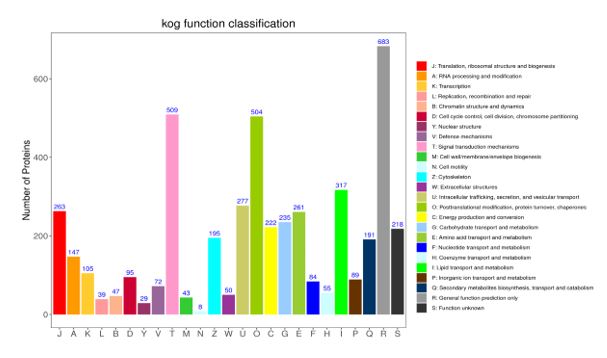
The identified proteins were compared with the KOG database, and the protein function was predicted and its function was classified and counted. A total of 3505 proteins were enriched. The functional annotation results show that these proteins are classified into 25 functional categories. (Figure 4). The annotations in the KOG database cover a wide range of protein functions, and we utilized it to perform statistics on the top 9 enriched functions. (Table 3). Among them, the top three representative functions were selected: functional prediction pathways, with a maximum enrichment of 638 proteins, Signal transduction pathway has an enrichment of 509 proteins and post-translational modification, protein turnover, and chaperones, which have an enrichment of 504 proteins.
Transcription Factor Analysis
Transcription factor (TF) is a protein molecule that can specifically bind to a specific sequence upstream of the 5'-end of a gene, thereby ensuring that the target gene is expressed with a specific intensity at a specific time and space. Due to the special significance of transcription factors, these proteins will be annotated and analyzed in depth0. The Plant TFDB (Plant Transcription Factor Database) and Animal TFDB (Animal Transcription Factor Database) contain the transcription factors and transcription factor family’s information of plants and animals respectively, and can predict whether the target protein is a transcription factor and the transcription factor family to which it belongs.
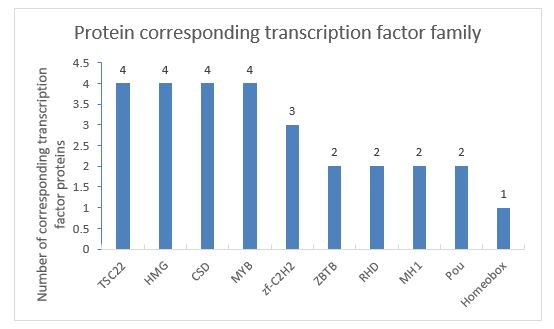
The analyzed species the Z. morio larvae is an animal, and the predicted protein sequences were compared with the corresponding TF database (plant TFdb/animal TFdb) by hmmscan. The database screened the transcription factors of the measured the Z. morio larvae proteins, and the results showed that 36 proteins belonged to the transcription factor category. The top 10 transcription factor families were counted, and the statistical results are as follows (Figure 5).
It can be seen from the statistical chart that the transcription factors TSC22, HMG, CSD, and MYB families are tied for first place in the protein of the Z. morio larvae, and each family accounts for 14.28% of the total matching proteins. The Homeobox family accounts for the least, with only 3.57%.
Identification and annotation result analysis
The protein results were identified by comparing the GO, KEGG, KOG databases, and the protein mainly participate in cellular processes, constitute cellular components, and perform binding functions. Among them, the proteins annotated by KEGG pathways are most associated with human diseases in terms of functional categories. However, this does not mean that the consumption of the protein can cause illness in humans. It can only indicate that the immune regulatory proteins of the Z. morio larvae are the majority among the enriched functional proteins. The KOG database has the highest number of enriched proteins, with a total of 3505, and is also the most comprehensive. Among them, functions of proteins related to nutrition, such as lipid transport and metabolism, amino acid transport and metabolism, carbohydrate transport and metabolism, and energy production and conversion, were all enriched in the top 9 functions. This indicates that the Z. morio larvae have better nutrient conversion and storage capabilities, reflecting their higher nutritional value, and the Z. morio larvae are easy to cultivate due to their vigorous food digestion and absorption0.
Discussion
As a new food, Z. morio larvae have been the focus of research in recent years. In this paper, 4,589 proteins were detected and identified by using DIA technology to construct GO, KEGG and KOG databases, and the annotation rate was 91.98%. Among them, protein functions related to nutrition, such as lipid transport and metabolism, amino acid transport and metabolism, carbohydrate transport and metabolism, energy production and conversion, are all concentrated in the first 9 functions of KOG annotation, which can predict that the nutritional value of Z. morio larvae is high. KEGG notes that the functional proteins of pathway are related to human diseases at most, and Trx-2 protein is found, which is related to immune regulation and reducing sulfur-oxygen conjugate. This study showed that the protein content of Z. morio larvae reached 51% (dry weight), which was significantly higher than that of Coleoptera related species Monochamus alternatus (23.99%) and Batocera horsfieldi (16.90%-26.54%), and close to that of water beetle (47.86%-48.74%) and Chestnut beetle (48%)[12].Compared with Lepidoptera insects (such as tussah pupa, bamboo worm, etc.), Z. morio is slightly lower than tussah pupa (60%) and oriental migratory locust (75%)[13-14], but it’s essential amino acids account for 36.53%, and its essential amino acid index is more than 92%, which is significantly higher than that of Monochamus alternatus (39.51%) and most traditional protein sources (such as beef) . In addition, the content of taste-presenting amino acids (glutamic acid, aspartic acid and glycine) of Z. morio is outstanding, which can improve the palatability of food processing [16]. Compared with Tenebrio molitor (essential amino acids account for 42.21%-44.01%), the amino acid balance of Z. morio is better [17].
The fat content of the Z. morio reached 32.70%, which was significantly higher than that of the Massicus raddei Blessig (13%) and the Batocera horsfieldi (2.20%-4.41%), but lower than that of water beetle [12]. Its fatty acids are mainly unsaturated fatty acids (such as linoleic acid and linolenic acid), which have the functions of regulating blood lipid and anti-inflammatory [15]. At the same time, the Z. morio is rich in trace elements such as Zn, Fe and Cu, and the content of heavy metals meets the food safety standards [18]. The synergistic effect of iron and protein may enhance its bioavailability, which is consistent with the research results of cricket powder [19]. Compared with bamboo worms (60% crude fat), golden cicada nymphs (9% fat) and other insects, Z. morio are more balanced in the ratio of "high protein to moderate fat", which meets the needs of modern functional food development.
In this study, Trx-2 protein was identified for the first time in Z. morio, which participated in antioxidant reaction through thioredoxin pathway. Unlike Lepidoptera insects (such as silkworm chrysalis) which rely on chitin to activate immunity, the discovery of Trx-2 protein in barley provides a new perspective for the immune regulation mechanism of Coleoptera insects [20]. In addition, the high expression of Trx-2 protein may explain the low oxidative stress of Z. morio, which has not been reported in water beetle [12]. Compared with plant-derived protein, the nutritional integrity of Z. morio protein is better, which may be related to its amino acid balance and fat-soluble vitamin content [21], providing a theoretical basis for developing anti-fatigue and antioxidant functional foods.
The protein quality (essential amino acid index > 92%) and the advantages of functional protein of Z. morio make it have broad prospects in the fields of feed replacement, biopharmaceuticals and so on. For example, compared with Tenebrio molitor, Z. morio has a more significant effect on improving muscle synthesis and endurance of exercise mice [21], suggesting its potential as a sports nutritional supplement. In addition, the antioxidant function of Trx-2 protein can be extended to the medical field, such as inflammatory regulation or adjuvant treatment of chronic diseases. In the future, it is necessary to further analyze the mechanism of Trx-2 and evaluate its stability under different processing conditions in order to promote industrial application. At present, the research on the metabolic pathway and functional protein interaction network of protein Group of Z. morio is insufficient, and its nutritional metabolism mechanism can be further analyzed by combining transcriptome and metabonomics. In addition, it is necessary to systematically evaluate the digestion and absorption characteristics of Z. morio protein, and compare it with feed insects such as housefly larvae (crude protein 33.3%-64.0%) and Black soldier fly (41.1%-43.6%) to clarify its application scenarios [18]. Optimizing amino acid composition or functional peptide segments through variety breeding will further enhance its market competitiveness.
- Wu F, Wu Z, Huang YT, et al. (2022) The effects of partial replacement of fishmeal with insect protein on the growth, muscle nutrition, and serum biochemical indices of mandarin fish (Siniperca chuatsi). Scientific Fish Farming, 37: 68-70.
- Rumbos CI, Athanassiou CG (2021) The superworm, Zophobas morio (Coleoptera:Tenebrionidae): a 'sleeping giant' in nutrient sources. Journal of Insect Science, 21: 1-11.
- Henry MA, Golomazou E, Asimaki A, et al. (2022) Partial dietary fishmeal replacement with full-fat or defatted superworm (Zophobas morio) larvae meals modulates the innate immune system of gilthead seabream, Sparus aurata. Aquaculture Reports, 27.
- Feng W (2023) Nutritional cognition, consumption behavior, and attitudes of urban and rural residents toward edible insects. Huazhong Agricultural University.
- Moreira NR, Cardoso C, Dias RO, et al. (2017) A physiologically-oriented transcriptomic analysis of the midgut of Tenebrio molito. Journal of Insect Physiology, 99: 58-66.
- Liu S, Rao XJ, Li MY, et al. (2015) Identification of candidate chemosensory genes in the antennal transcriptome of Tenebrio molitor (Coleoptera: Tenebrionidae). Comparative biochemistry and physiology. Part D, Genomics & proteomics, 13: 44-51.
- Zhao H, Zhao SG, Du YL, et al. (2023) Transcriptomic differences analysis of four subspecies of overwintering western honeybee. Acta Entomologica Sinica, 66: 381-90.
- Guo R, Chen DF, Huang ZJ, et al. (2017) Transcriptomic study of pathogens in the intestinal process of Chinese honeybee larvae under coccidious streess. Acta Microbiologica Sinica, 57: 1865-78.
- Wei XW, Zhang JF. Research progress of cellular particle proteomics. Medical Review, 22: 457-60.
- Guo HR, Li J, Zang Y (2023) Regulatory mechanism of transcription factors into the nucleus. Chinese Journal of Biochemistry and Molecular Biol, 39: 683-91.
- Wang XN (2012) Research on the biology and artificial breeding conditions of Zophobas morio. Sichuan Agricultural University.
- Ma YK, Zheng LK, Sun ZW, et al. (2002) Analysis of nutritional components in three aquatic beetles. Acta Nutrimenta Sinica, 25: 90-2.
- Liu ZY, Li L, Wu GH (2019) Overview of nutritional functions of several common edible insects. China Sericulture, 40: 48-51.
- Hu AX, You S (2024) Current status and prospects of research on edible insects. Science of Sericulture, 50: 469-75.
- Hlongwane ZT, Slotow R, Munyai TC (2020) Nutritional Composition of Edible Insects Consumed in Africa: A Systematic Review. Nutrients, 12.
- Zhu F, Shi ZH (2022) Resource characteristics and development prospects of edible insect protein. Journal of Chinese Institute of Food Science and Technology, 22: 44-52.
- Xu QA, Peng WL, Li XX, et al. (2008) Research progress on economic insects: tenebrio molitor and Zophobas morio. Anhui Agricultural Science Bulletin, 21: 158-60.
- Wang YD (2022) Research progress on the utilization of insects as feed and their impact on the ecological environment. Hunan Feed, 25: 17-22.
- Agbemafle I, Hanson N, Bries AE, et al. (2019) Solanum torvum alternative protein and iron sources from edible Insects but not improved body composition and iron status in malnourished rats. Nutrients, 11.
- Lan Y, Wang ZY, Zhao XM, et al. (2024) Research progress on the nutritional value and edible safety of edible insects. Journal of Food Safety and Quality, 15: 27-33.
- Shi L, Yang W, Yang CP, et al. (2015) Effects of Zophobas morio protein on muscle, blood indices, and exercise endurance in exercising mice. Science and Technology of Food Industry, 36: 359-62.

Tables at a glance
Figures at a glance